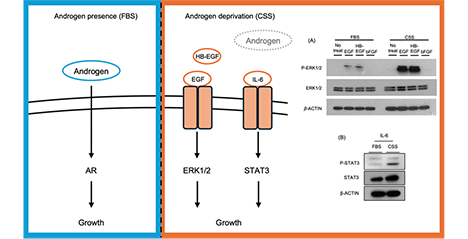
Prostate cancer grows in an androgen receptor (AR)-dependent manner; however, some cases progress to castration-resistant prostate cancer (CRPC) after androgen deprivation therapy. One of the molecular mechanisms underlying this transition is the activation of AR-independent “bypass signaling” pathways. Although several signaling pathways have been implicated, the mechanisms that regulate these signaling pathways remain incompletely understood. In this article, we focus on cell surface glycans, particularly sulfated glycosaminoglycans (GAGs), and summarize two independent mechanisms involved in the acquisition of castration resistance: activation of epidermal growth factor receptor (EGFR) signaling mediated by 3-O-sulfated heparan sulfate (3-OS HS) and enhancement of interleukin (IL)-6/signal transducer and activator of transcription (STAT)3 signaling mediated by chondroitin sulfate E (CS-E). These findings highlight the potential of GAGs as novel therapeutic targets in CRPC. ...and more
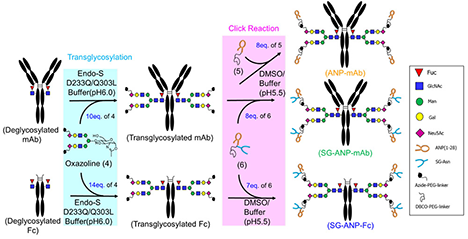
Glyco-engineering is an advanced technology that artificially designs and modifies the composition and structure of glycans bound to biomolecules such as peptides and proteins. In the biopharmaceutical field, a wide range of approaches have been investigated that aim to enhance functionality, enhance target specificity, and reduce side effects by modulating drug properties such as stability, efficacy, safety, and pharmacokinetics. Particularly, chemoenzymatic glycoengineering of monoclonal antibodies using endo-beta-N-acetylglycosaminidase (ENGase) is known to provide useful tools such as linkers for site-specific conjugation of drugs to antibodies. This review provides recent updates of drug discovery research using chemoenzymatic approaches and developments to overcome challenges in large-scale production. ...and more
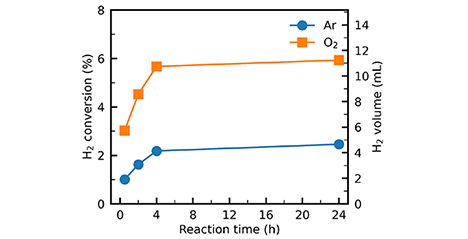
Although hydrogen energy has attracted considerable attention as a means of reducing dependence on fossil fuels and mitigating climate change, current hydrogen production relies predominantly on fossil-fuel-based processes. In this study, we discovered that mixing Japanese cedar (Cryptomeria japonica) wood powder with sodium hydroxide in a solid–solid state and heating the mixture at temperatures as low as 25–120°C is sufficient to generate hydrogen gas. This so-called "low-temperature reaction" proceeds via a reaction pathway entirely distinct from conventional thermochemical processes such as pyrolysis, and was confirmed to occur with all three major constituents of wood. The findings suggest promising applications in utilizing industrial waste heat and enabling low-cost, small-scale hydrogen production. ...and more
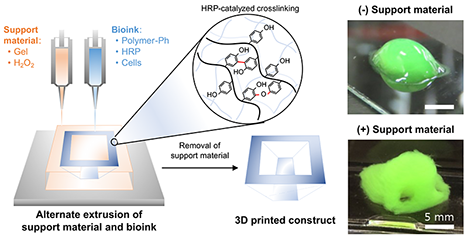
In the fields of regenerative medicine, drug discovery, and disease modeling, three-dimensional (3D) bioprinting has attracted considerable attention as a technology for arranging cells in three dimensions and reconstructing tissue-specific microenvironments. In this approach, living cell-containing inks (bioinks) are used to fabricate 3D structures. Conventional tissue engineering has generally relied on seeding cells onto prefabricated scaffolds, whereas 3D bioprinting enables the simultaneous placement of cells and materials at desired locations during construct formation, thereby allowing the fabrication of more complex architectures that better resemble native tissues. ...and more
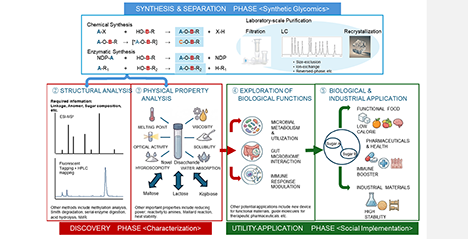
Disaccharides, the simplest oligosaccharides, can theoretically form up to 3,056 unique structures from various D- and L-aldohexose combinations of two monosaccharide units. While most of these have not been studied, advances in rare sugar synthesis now allow their experimental investigation (Nakakita SI, Hirabayashi J, BBA Advances, 7:100143, 2025). This opens new possibilities for discovering disaccharides with unique properties and potential uses as sweeteners, pharmaceuticals, and biodegradable materials. Compared to more complex oligosaccharides, disaccharides are easier to study systematically. We propose “disaccharide glycomics” as a systematic framework to explore this newly accessible chemical space, opening the door to discoveries that are both scientifically and practically significant. ...and more