
Tadasu Urashima
After graduation from Tohoku University (Doctor of Agriculture) in 1986, he started his professional career by studying milk oligosaccharides at Obihiro University. In 1991, he studied glycosyltransferase activity in lactating mammary glands of the tammar wallaby (marsupial) under Dr. Michael Messer in the Department of Biochemistry, the University of Sydney. Then he completed a comprehensive study of milk oligosaccharides of monotremes, marsupials, and several species of eutherians with Dr. Messer. He is interested in how the present milk components have been acquired during the evolution of mammals, especially how the acquisition of α-lactalbumin, a milk protein, has resulted in the appearance of milk oligosaccharides and lactose, and caused their biological significance to change during evolution. In 2003 ~ 2022, he had been a full professor at Obihiro University, and is an honorary professor today. At present he is president of the Japanese Society of Dairy Science and Japanese Consortium for Glycoscience and Glycotechnology (JCGG).

Jun Hirabayashi
Tokai National Higher Education and Research System, Nagoya University, Japan. Ph.D, Science.
After graduating from Tohoku University (Master of Science), he started his professional career at Teikyo University under the supervision of Prof. Kenichi Kasai to investigate animal lectins. On the occasion of GlycoXV (Tokyo, 1999), he proposed the concept of glycome; for this structural aspect, he moved to the National Institute of Advanced Industrial Science and Technology (AIST, Tsukuba) in 2002, and was involved in a series of national projects for glycan engineering, while he was a deputy director in Research Center for Medical Glycoscience (2006~), and a prime senior researcher of Research Center for Stem Cell Engineering (2012~). Now, he is a designated professor at the Institute for Glyco-core Research (iGCORE), Tokai National Higher Education and Research System, Nagoya University, and a project manager (human glycome). At iGCORE, he is pleased to witness the final phase of realization of the concept of glycome, which he initially proposed over a quarter century ago. He is a vice president of the Japanese Society of Carbohydrate Research (JSCR) and Japanese Consortium for Glycoscience and Glycotechnology (JCGG), as well as a visiting professor of both Kagawa University (2003~) and Yokohama City University (2019~).
Human milk contains 12~13 g/L of oligosaccharides, of which around 250 varieties have been separated and more than 160 structures have been characterized1-3. Human milk oligosaccharides (HMOs) have been classified into 20 series based on the core structures, which contain neither fucose nor sialic acid, and the biosynthetic pathways of these structures have been proposed based on the activities of β-1-3-N-acetylglucosaminyltransferase (iGnT), β-1-6-N-acetylglucosaminyltransferase (IGnT), β-1-3-galactosyltransferase (β3GalT) as well as β-1-4-galactosyltransferase (β4GalT)2,3. Around 50 varieties of oligosaccharides have been also separated and characterized from bovine colostrum4-7, and the biosynthetic pathways of their core structures have been proposed as well8,9.
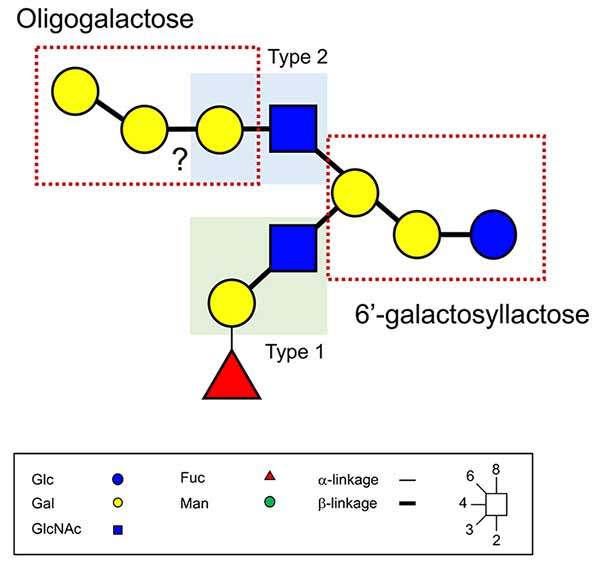
Recently, however, a significant number of novel milk oligosaccharides have been reported, whose biosynthesis cannot easily be understood based on the previously proposed mechanisms. Hanisch and Kunz (2021) characterized a new HMO structure (Fig. 1)10: it consists of both type 1 (Galβ1-3GlcNAc) and type 2 (Galβ1-4GlcNAc) lactosamine structures linked to the same non-reducing terminal Gal residue of the reducing trisaccharide 6’-galactosyllactose (6’-GL). On the other hand, at the non-reducing ends, an H antigen (Fucα1-2Gal) and polygalactose structures are attached to the β1-3 and β1-6 GlcNAc branches, respectively (Fig. 1)10. More recently, the group of Dr. C.K. Ni of Taiwan Academia Sinica reported a series of novel and quite amazing oligosaccharide structures from the milks of humans, cows, and goats11. After extraction of the carbohydrates from the milks, fractions containing oligosaccharides were separated by size exclusion chromatography and high-performance liquid chromatography (HPLC) on an Amide-80 column, then separated by HPLC using a porous graphite carbon column (PGC), and finally analyzed by a new mass-spectrometric analysis method, named logically derived sequence (LODES) multistep tandem mass spectrometry (LODES/MSn), to determine oligosaccharide structures.
Although it was not easy to determine the component monosaccharides in terms of diastereomers, anomeric configuration of the glycosidic linkage, and the linked position of each monosaccharide residue in the oligosaccharide structure by mass spectrometry, his group found the empirical rules to identify the structures based on the fragmentation patterns that had been established with standard saccharides by LODES/MSn12,13. The principles underlying the rules are as follows: (1) the dehydration reaction mainly occurs at the reducing end of the sodium or lithium adducts of oligosaccharides during the fragmentation, and (2) the ring cleavage by the retro aldol reaction mainly occurs at the reducing end; these fragmentation patterns are useful to determine the linkage position of the reducing end residue of each saccharide14-16. The reducing residues were labeled with 18O before analysis. Once the structures were established by LODES/MSn, corresponding standard oligosaccharides were chemically synthesized, and their retention times as well as the fragmentation patterns were compared by means of HPLC and mass spectrometry analyses. Upon separation by PGC-HPLC, no saccharide was subjected to reduction so that both α and β anomers at the reducing end were maintained; hence their retention times as well as the fragmentation patterns in the MS analysis were compared between the milk oligosaccharides and the corresponding authentic standards. The tri- and tetrasaccharides thus characterized were classified as shown in Fig. 2 based on the hypothesized biosynthetic pathways (discussed below). Although these structures are not readily acceptable, we must believe that they are correct considering the analysis was precisely designed and performed consistently. These novel structures contain neither fucose nor sialic acid, but are assumed to be novel combinations of glucose, galactose, N-acetylglucosamine as well as N-acetylgalactosamine.
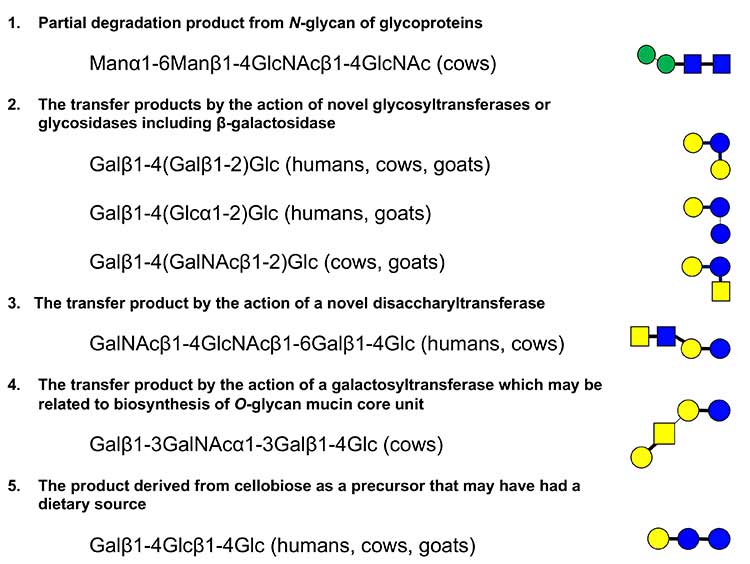

Among the classified categories, the first is potential degradation products such as Manα1-6Manβ1-4GlcNAcβ1-4GlcNAc, which should be produced from a class of N-glycans by the action of some endoglycosidases. It is also possible that they are produced from some intermediate components linked to dolichol phosphate, which may be produced during the biosynthesis of N-glycans by the action of phosphatase. The second category consists of branched type trisaccharides of Galβ1-4(Galβ1-2)Glc, Galβ1-4(Glcα1-2)Glc, and Galβ1-4(GalNAcβ1-2)Glc; in all oligosaccharides of this category, the reducing terminal glucose is 1-2 branched. They might be the reaction products of the action of unknown branched type glycosyltransferases. Alternatively, it is also possible to speculate that Galβ1-4(Galβ1-2)Glc as well as the previously identified galactosyllactoses, i.e., Galβ1-6Galβ1-4Glc (6’-GL) and Galβ1-3Galβ1-4Glc (3’-GL), are produced by β-galactosidase working in transglycosylation mode in mammary epithelial cells (Note 1). In fact, the commercial galacto oligosaccharides with lactose are manufactured using β-galactosidase17,18. In a similar manner, Galβ1-4(Glcα1-2)Glc and Galβ1-4(GalNAcβ1-2)Glc might be the reaction products of α-glucosidase and β-N-acetylgalactosaminidase, respectively, but not glycosyltransferases (Note 2). In this case, the enzymes transfer glucose or N-acetylgalactosamine to lactose, respectively.
Note 1: β-galactosidase can be used for not only hydrolysis but also transgalactosylation when the concentration of lactose as substrate is high in the reaction mixture (note that transglycosylation is not a reverse reaction of hydrolysis). This reaction can proceed as follows: Galβ1-4Glc + Galβ1-4Glc → Galβ1-6Galβ1-4Glc + Glc. The commercial galacto oligosaccharides are a mixture of reaction products manufactured by the same process18; they are Galβ1-6Gal, Galβ1-3Gal, Galβ1-3Glc, Galβ1-2Glc, Galβ1-6Glc, Galβ1-4(Galβ1-6)Glc, Galβ1-6Galβ1-6Glc, Galβ1-6Galβ1-4Glc (6’-GL), Galβ1-4Galβ1-4Glc (4’-GL), Galβ1-3Galβ1-4Glc (3’-GL), Galβ1-6Galβ1-6Galβ1-4Glc, Galβ1-3Galβ1-3Galβ1-4Glc, and Galβ1-6Galβ1-6Galβ1-6Galβ1-4Glc. It is assumed that some disaccharides are hydrolysis products from trisaccharides. The disaccharide Galβ1-2Glc is possibly derived from Galβ1-4(Galβ1-2)Glc by hydrolysis. It is possible to obtain a different mixture of galacto oligosaccharides by using other β-galactosidases of different specificities.
Note 2: In general, the linkage specificity of glycosyltransferases is strictly controlled, i.e., the position of the monosaccharide transferred to an acceptor saccharide is rigorously specific, while that of glycosidases is far less specific. Therefore, resultant products are sometimes unusual (or unnatural) in the reaction of glycosidases. For example, Galβ1-4Glcβ1-1αFuc was found as a product, when the reaction was designed using lactose and fucose as substrates with α-fucosidase from the fungus Aspergillus niger19. It is not easy to imagine that Galβ1-4(Glcα1-2)Glc or Galβ1-4(GalNAcβ1-2)Glc would be biosynthesized by the usual glycosyltransferases. Rather, they might have been produced by the transfer reaction of Glc or GalNAc to lactose by mammary gland α-glucosidase or βN-acetylgalactosaminidase, respectively.
The potential biosynthetic pathway of GalNAcβ1-4GlcNAcβ1-6Galβ1-4Glc is classified as the third category. Weng et al.11 suggested that this tetrasaccharide is formed by the direct transfer of the disaccharide, GalNAcβ1-4GlcNAc (LacdiNAc), to lactose, because GlcNAcβ1-6Galβ1-4Glc was not found in any of the human and bovine milk samples used in the study (i.e., if this trisaccharide is found in milk, one would imagine that the tetrasaccharide is synthesized from the trisaccharide as a precursor by the action of β4N-acetylgalactosaminyltransferase). For the transfer of the disaccharide to occur, however, GalNAcβ1-4GlcNAc-UDP must be present as a prerequisite donor substrate of this novel glycosyltransferase (Note 3).
Note 3: As described below, UDP di- or trisaccharides have been identified in some milks/colostra, however, their existence has never confirmed in a consistent manner. While it is likely that a still undiscovered, but major biosynthetic pathway is used to produce these compounds, it is possible that these products are the results of errors in the synthetic process. For example, one can speculate that when GalNAcβ1-4GlcNAc-UDP is synthesized (if this component is present), the synthetase for GlcNAc-UDP catalyzes transferring UDP to GalNAcβ1-4GlcNAc in place of GlcNAc, an original substrate, by mis-reaction.
Thus, the following reactions are assumed:
1) original reaction: GlcNAc-P + UTP => GlcNAc-UDP +PP
2) mis-reaction: GalNAcβ1-4GlcNAc-P + UTP => GalNAcβ1-4GlcNAc-UDP + PP
Namely, in the reaction 2, the enzyme binds to and acts on the wrong substrate GalNAcβ1-4GlcNAc-P in place of GlcNAc-P to produce GalNAcβ1-4GlcNAc-UDP.
There is another potential reaction leading to the biosynthesis of GalNAcβ1-4GlcNAc-UDP. Namely, β4GalNAc transferase, which originally transfers GalNAc from GalNAc-UDP to GlcNAc-R, mis-reacts with GlcNAc-UDP as an acceptor and transfers GalNAc from GalNAc-UDP to GlcNAc-UDP.
3) original reaction: GalNAc-UDP (D) + GlcNAc-R (A) + => GalNAcβ1-4GlcNAc-R
4) mis-reaction: GalNAc-UDP (D) + GlcNAc-UDP (A) + => GalNAcβ1-4GlcNAc-UDP
In this case, the reaction 4) occurs in place of 3) and is the result of mis-reaction with GalNAc-UDP (D).
We designate this type of mis-reaction as “molecular wobbling” of the enzyme. In the “molecular wobbling” type mis-reaction, the original acceptors, as well as donors, are altered under some conditions leading to modified reactions. In the spin-off version of the series Galectins (Glycoforum. 2021 Vol.24(3), A6), we described that the acceptor for β4Gal-T1 is switched from GlcNAc to Glc by the association with the modifier milk protein α-lactalbumin as a representative modifier20. In this case, α-lactalbumin is necessary for this wobbling to occur. Could the GlcNAc-UDP synthetase associate with some modifier protein like α-lactalbumin, if the wobbling really occurs? At this moment, this is just a hypothesis.
Galβ1-4GlcNAc-UDP but not GalNAcβ1-4GlcNAc-UDP has been found in the milks/colostra of humans21, pigs22 and reindeer23. Previously, Dr. Akira Kobata identified Galβ1-4GlcNAc-UDP21 and Fucα1-2Galβ1-4GlcNAc-UDP24 in human milk. Then he attempted, in vain, to find relevant glycosyltransferases that would transfer such di- or trisaccharides to appropriate acceptors20. As described in a spin-off version of “Milk oligosaccharides and galectins” in the Glycoforum series “Galectins”20, UDP-oligosaccharides are sometimes found in the milks/colostra of humans21, 24, goats25, and sheep26, although their consistent existence remains to be proven27. Weng et al.11 also hypothesized that the biosynthesis of Galβ1-4GlcNAcβ1-3Galβ1-4Glc (LNnT) and Galβ1-4GlcNAcβ1-6Galβ1-4Glc (iso LNnT) in bovine milk, which are not novel oligosaccharides, might have occurred by block transfer of the disaccharide (Galβ1-4GlcNAc) to lactose, because the trisaccharides, GlcNAcβ1-3Galβ1-4Glc and GlcNAcβ1-6Galβ1-4Glc, were not found in the bovine milk. Even though attempts to identify such an oligosaccharyltransferase have never succeeded, it would be exciting if someone found the activity of this type of enzyme. In that case, one may also hypothesize that (i/I) N-acetylglucosaminyltransferase uses not only UDP-GlcNAc but also Galβ1-4GlcNAc-UDP as donors. Although Galβ1-4GlcNAc-UDP has never been identified in bovine milk/colostrum, one could suppose its existence in this resource, as it exists in those of humans21, pigs22 and reindeer23 (Note 4).
Note 4: Regarding the block-transfer reaction with di- or trisaccharide units in the biosynthesis of glycoconjugates, one can speculate the presence of two potential synthetases. One is a novel oligosaccharyltransferase, which has the substrate specificity for UDP di- or trisaccharides, and the other is any of the known glycosyltransferases, which originally transfer monosaccharide from UDP monosaccharide, but then transfer di- or trisaccharide from UDP di- or trisaccharide by a molecular wobbling mechanism. Although there have been some attempts to find oligosaccharyltransferase, none have succeeded as described above. In the latter possibility, one can assume that molecular wobbling enables GlcNAc transferase to use Galβ1-4GlcNAc-UDP, for example, as a donor substrate in place of the original donor GlcNAc-UDP.
Thus, the following reactions are assumed:
5) original reaction: GlcNAc-UDP + Gal-R → GlcNAcβ1-3Gal-R
6) mis-reaction: Galβ1-4GlcNAc-UDP + Gal-R → Galβ1-4GlcNAcβ1-3Gal-R
In this case, reaction 6), but not 5), occurs. However, does GlcNAc transferase need to associate with some protein for wobbling of this enzyme to occur as described above? One may also speculate that this mis-reaction by molecular wobbling due to the rather ambiguous substrate specificity of the enzyme occurs under a different set of pH or substrate concentration conditions, unless modifier proteins like α-lactalbumin intervene. A clear molecular basis for the modification of acceptor specificity in the case of α-lactalbumin with β4Gal-T1 has been established. We may define reactions as “wobbling” reactions when the molecular basis is clear and as mis-reaction when the basis is unclear. Further discussion is needed about the proper terminology to use for these unexpected reactions.
Although found at the non-reducing end unit of the N-glycan or as a free disaccharide in bovine milk/colostrum4, 28,29, GalNAcβ1-4GlcNAc (LacdiNAc) has never been found in human milk. The finding of the tetrasaccharide containing non-reducing LacdiNAc units in human milk is interesting from the perspective of novel HMO biosynthesis.
The explanation took a while for the 3rd category, but the 4th category is Galβ1-3GalNAcα1-3Galβ1-4Glc, which was identified in bovine milk, can be biosynthesized from precursor GalNAcα1-3Galβ1-4Glc by transgalactosylation. GalNAcα1-3Galβ1-4Glc has been identified in the milks/colostra of goats and sheep4,5 as well as cows4,5,30. Incidentally, Galβ1-3GalNAc, the non-reducing disaccharide unit of this tetrasaccharide is common to the core 1 structure of mucin type O-glycan, also known as T antigen31. Therefore, it may be speculated that Galβ1-3GalNAcα1-3Galβ1-4Glc is also synthesized from GalNAcα1-3Galβ1-4Glc by the action of β3galactosyltransferase, which also produces the T antigen.
As Weng et al.11 pointed out the most amazing of Galβ1-4Glcβ1-4Glc in the milks of humans, cows, and goats (classified as the 5th category), it is a widely accepted rule that milk oligosaccharides including HMOs are biosynthesized from lactose as a starting point (of note, a few oligosaccharides found in the milks/colostra of cows and goats contain the reducing unit Galβ1-4GlcNAc). However, the reducing disaccharide unit of this trisaccharide is Glcβ1-4Glc (cellobiose), not lactose. Although it is possible to assume that the trisaccharide is biosynthesized by the cooperative action of β4GalT-1 with α-lactalbumin, which transfers galactose to cellobiose, it is difficult to imagine that cellobiose could have been biosynthesized within mammalian body tissues including mammary glands. Could dietary cellobiose be transported into the mammary gland epithelial cells after its absorbance in the small intestine and successive internal circulation? Another possibility is that the disaccharide, Galβ1-4Glc, is bulk-transferred to glucose. Could there be an unknown enzyme, which transfers the disaccharide block from Galβ1-4Glc-UDP as a donor to glucose as an acceptor? Alternatively, all of these products described here are results of wobbling by known enzymes, e.g., glycosyltransferases, glycosidases, sugar-nucleotide syntheses. Anyway, this represents a new puzzle.
It has been known that the concentration of galactosyllactose (3’-GL and 6’-GL) in human milk is lower by two orders of magnitude than those of major oligosaccharides, such as 2’-FL (Fucα1-2Galβ1-4Glc) and LNT (Galβ1-3GlcNAcβ1-3Galβ1-4Glc)32. However, the concentrations of the novel trisaccharides described above (the second category) are lower still than those of 3’-GL and 6’-GL in human milk. Although the concentrations of the novel trisaccharides are also low in bovine and caprine milks, their concentrations are rather comparable to those of 3’-GL, 6’-GL, and Galα1-3Galβ1-4Glc (isoglobotriose), and thus, there is no great difference among them, which is distinct from the human milk.
What is the biological significance of the novel tri- and tetrasaccharides? No doubt, the development of the new analytical techniques should bring us to a new frontier in chemistry of milk oligosaccharides. It is hoped that in addition to their confirmation, the present experimental results will stimulate further discussion about the biological functions of milk oligosaccharides including the new structures investigated in this study.