
Shou Takashima
Researcher, Team leader, Laboratory of Glycobiology, The Noguchi Institute
Dr. Shou Takashima received his Ph.D. in 1997 from the Graduate School of Agricultural and Life Sciences, University of Tokyo. He worked as a postdoctoral fellow at RIKEN and Fukushima Medical University. He has been working at the Noguchi Institute since 2008. He has been studying carbohydrate-related enzymes such as sialyltransferases, cellulases, endo-β-N-acetylglucosaminidases, O-glycan related enzymes, etc.
Glycans play important roles in protein function, stability, intermolecular interaction, and so on. These are highly related to glycan structures in many cases. However, the structure of the glycan attached to the specific glycosylation site usually varies in each molecule of glycoprotein, and this variation gives rise to the heterogeneity and functional differences between glycoproteins. Therefore, glycan remodeling aims to increase the homogeneity and strengthen the function and stability of the glycoprotein by attaching glycans of specific structure to the glycosylation sites of the glycoprotein. The glycan remodeling techniques have been widely developed, and one of them is the endo-β-N-acetylglucosaminidase (ENGase, EC 3.2.1.96)-based method. In connection with this, this review outlines the basic knowledge of glycan remodeling and the recently identified novel ENGases that recognize the non-reducing end of N-glycans and can hydrolyze multibranched complex type N-glycans.
Glycans attached to proteins are roughly divided into two groups. One is N-glycans, which attach to the Asn residue in the consensus sequon Asn-Xaa-Ser/Thr (Xaa ≠ Pro) in a protein. The other is O-glycans, which attach to the Ser/Thr residues in a protein. Glycan remodeling with ENGases applies to only the remodeling of N-glycans. ENGases are enzymes that release N-glycans by hydrolyzing the bond between two N-acetylglucosamine (GlcNAc) residues located at the reducing end of the N-glycan. Among ENGases, there are enzymes that catalyze the transglycosylation reaction, which is the reverse reaction of the above hydrolysis reaction. After the hydrolysis reaction by ENGase, one GlcNAc residue remains on a protein or a peptide, where the N-glycan of the specific structure is transferred by the transglycosylation activity of ENGase. This is the basic principle of the glycan remodeling with ENGases1-3. In this method, highly homogenous glycoproteins can be obtained when using chemically synthesized high-grade N-glycan.
Other techniques are available for glycan (O-glycan as well as N-glycan) remodeling, for example, the method for producing the target glycoprotein in cells with suppressed or overexpressed specific glycogene expression4-6, or the method for creating the target glycan on the protein by using glycosylhydrolases (sialidase, galactosidase, etc.) and/or glycosyltransferases (GlcNAc transferase, galactose [Gal] transferase, sialyltransferase, etc.) for remodeling7.
According to the classification by the Carbohydrate-Active enZYmes (CAZy) database8,9, ENGases are classified into two families, Glycoside hydrolase (GH) 18 and GH85. Besides ENGases, chitinases are included in the GH18 family. Among GH18 and GH85 ENGases, the amino acid sequence motifs, DxxDxDxE and NxE, are conserved in their active sites, respectively. The ENGase mutants called “glycosynthases,” whose above conserved motifs are modified for suppression of hydrolysis activity and expression of transglycosylation activity, are utilized in the transglycosylation reaction of glycan remodeling with ENGases.
N-glycans are roughly divided into three groups (Fig. 1). The high mannose (Man) type N-glycans are composed of Man residues except for two GlcNAc residues located at the reducing end of N-glycans. The complex type N-glycans have two GlcNAcβ1,2Man structures and include sugar components such as sialic acid (Sia), Gal, and fucose (Fuc) besides Man and GlcNAc. They also include multibranched structures, and/or α1,6-linked Fuc attached to the GlcNAc residue at the reducing end, which is called the “core Fuc.” Thus, the structures of complex type N-glycans are diverse. Hybrid type N-glycans have both the high Man type N-glycan branch and the complex type N-glycan branch with a GlcNAcβ1,2Man structure. All these N-glycans have the common core structure (Man3GlcNAc2) at the reducing end.
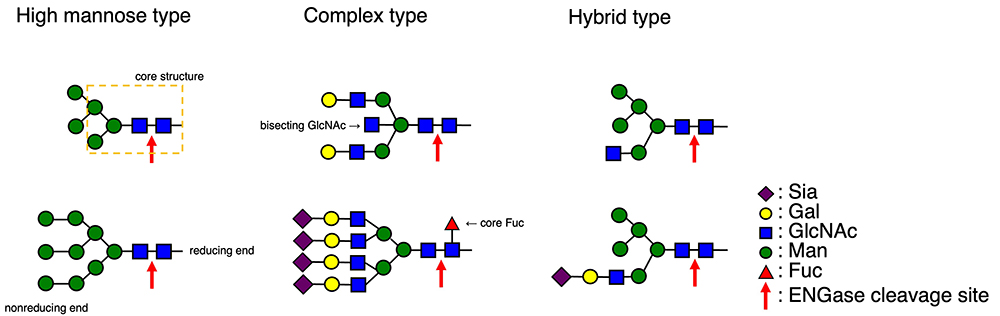
Many ENGases show substrate specificity toward these N-glycans1,2. In the case of glycan remodeling with ENGases, N-glycans on the target protein must first be removed by ENGase, which results in the acceptor substrate with one GlcNAc residue (+ core Fuc) (Fig. 2). Therefore, the choice of a suitable ENGase depends on which types of N-glycans are on the target protein. Among existing ENGases10, enzymes such as Endo H, which hydrolyzes high Man type and hybrid type N-glycans, Endo-M, which hydrolyzes non-corefucosylated bi-antennary complex type N-glycans (the W251N mutant, which can hydrolyze corefucosylated bi-antennary complex type N-glycans, is also available11), and Endo F3, which prefers to hydrolyze corefucosylated complex type N-glycans, are commercially available. As for the hydrolysis of N-glycans on IgG antibodies, enzymes such as Endo S and Endo S2, which exhibit the high specificity toward N-glycans on the Fc domain of IgG antibodies, are used preferentially.
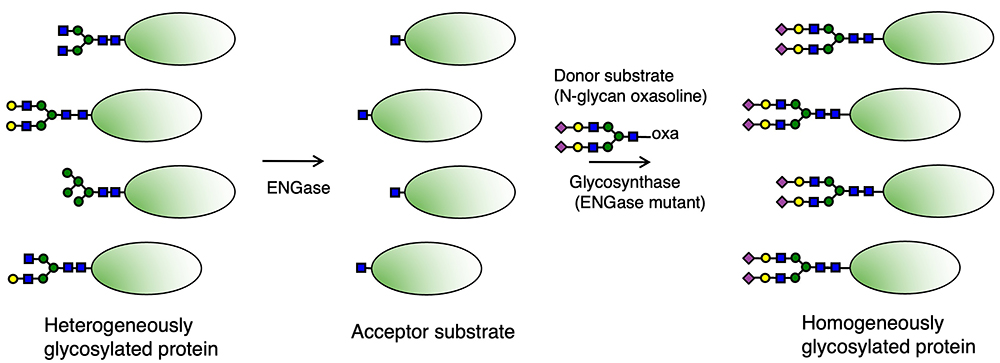
As described above, complex type N-glycans include glycans with branched structures such as tri-, tetra-, and penta-antennary and bisecting GlcNAc structures. As for the complex type N-glycans with tri- and tetra-antennary structures and/or bisecting GlcNAc structure, these are ubiquitously observed in many human glycoproteins, and the penta-antennary N-glycans are observed in glycoproteins from birds and fishes12,13. When preparing the acceptor substrate from the glycoprotein with such complicated branched complex type glycans, it is necessary to remove these glycans completely by ENGase. Otherwise, it is difficult to obtain the homogeneously glycosylated glycoprotein by glycan remodeling with ENGases, as some originally attached glycans may remain on the acceptor substrate.
So far, the reports of ENGases that can efficiently hydrolyze multibranched complex type N-glycans such as tetra-antennary glycans have been limited14-16. However, novel ENGases showing high hydrolysis activity toward multibranched N-glycans with or without core Fuc have been identified recently. Among them, there are GH85 family ENGases from Tannerella species (Endo-Tsp1006, Endo-Tsp1263)17. These enzymes have characteristic features: they recognize the nonreducing ends of glycans and selectively hydrolyze multibranched complex type glycans. Namely, Endo-Tsp1006 and Endo-Tsp1263 preferentially hydrolyze complex type N-glycans with α2,6-linked Sia or Gal residues and GlcNAc residues at the nonreducing end of glycans, respectively (Fig. 3). No other ENGases are known to have such features, and it may be possible for these enzymes to hydrolyze or retain only the glycans of specific structure on the glycoprotein. At present, by using the novel ENGases including Endo-Tsp1006, Endo-Tsp1263, and their homolog enzymes14,15,18,19, acceptor substrates for the glycan remodeling with ENGases can be prepared from various glycoproteins.
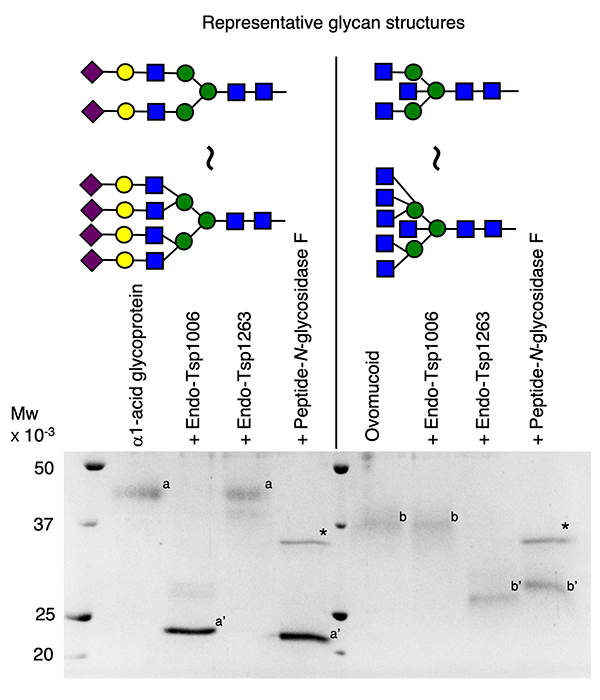
In glycan remodeling with ENGases, the glycan is transferred to the acceptor substrate by an enzyme such as an ENGase mutant. However, ENGase mutants that can transfer multibranched N-glycans of more than tetra-antennary glycans efficiently to the glycoprotein acceptor have not yet been developed. In the case of the above-described Endo-Tsp1006, Endo-Tsp1263, and their homolog enzymes, transglycosylation activity toward small molecules such as monosaccharides has been observed16, but not for glycoprotein acceptors.
Although the glycan remodeling with ENGases has been applied to the glycoengineering of antibody and some glycoproteins, the scope of their application has been limited. However, once they become available, ENGases that can hydrolyze and transfer various glycans including multibranched complex type glycans will be powerful tools for the glycan remodeling. It is expected that the development of such enzymes will proceed and the scope of application for glycan remodeling with ENGases will expand.