
Toshinori Sato
Current position: Professor at the Department of Biosciences & Informatics, Faculty of Science and Technology, Keio University
Academic background: Dr. Sato completed his master’s course at the Kyushu University Graduate School of Engineering Sciences in March 1983, withdrew from his doctoral course in September 1983, and obtained a doctoral degree in engineering from Kyoto University in 1990.
Work career: He was appointed Assistant Professor at the Nagasaki University Faculty of Engineering in October 1983, Assistant Professor at the Kyoto University Faculty of Engineering in October 1990, Associate Professor at the Tokyo Institute of Technology Faculty of Life Science and Technology in April 1992, and Associate Professor at Keio University Faculty of Science and Technology in April 2000. He has held his current position since April 2002.
Dr. Sato was a recipient of the CSJ Award for Young Chemists in 1993, and the Bio Business Competition Japan The Takeda Foundation Award in 2010.
Research interests: Dr. Sato’s research interests include glycomics using saccharide primers, polysaccharide-based drug delivery systems, lipid raft formation and recognition functional analysis using biomembrane models, and search and functional analysis of glycan-related peptides using a phage-display method.
Saccharide primers are sugar derivatives that serve as precursors in the glycan biosynthetic pathway. Saccharide primers can interact with cells to produce glycosylated products via the glycan biosynthetic pathway. The sequence of the glycosylated product can be analyzed by liquid chromatography - mass spectrometry (LC-MS) for glycan profiling and library construction. This paper introduces a saccharide primer method as a novel tool of glycomics.
Glycans expressed in cells are presented on cell surfaces as a wide variety of glycoconjugates such as glycolipids, glycoproteins, and proteoglycans, and are closely related to various cell functions such as genesis, differentiation, proliferation, and migration. To understand cell functions, it is necessary to understand the detailed structure of the glycans expressed in the cell. To this end, there is a demand for glycan profiling. Expression analyses of genes and proteins are carried out actively, and genomics and proteomics have developed dramatically. In the meantime, glycan analysis has not been established as a common technology in the life sciences fields. For this reason, there has been no spread of research in glycomics. To overcome this situation and to advance glycan research, research and development of convenient methods of glycan analysis is indispensable.
There are two main aims of glycomics: the first is to clarify the detailed structure of the glycans expressed in cells, and the second is to elucidate the role of the glycans. If the two are achieved, it will be possible to make the scientific significance of glycan functions in the life sciences known to many researchers, and thereby to possibly contribute to the diagnosis of diseases and to drug discovery. To attain the goals of glycomics, glycoscience for identification of glycans expressed in cells and elucidation of their functions, and glycotechnology for preparing glycans and their derivatives and verifying their functions are both indispensable. Since glycoscience and glycotechnology each require levels of high expertise, they have been performed and developed by different researchers; however, reformative development has been underway through collaboration of researchers in the two fields. In this situation, the author has been engaged in the development of a saccharide primer method as a technology for bridging glycoscience and glycotechnology. This method is outlined below.
A saccharide primer is a precursor for the biosynthesis of glycans as substrates of glycotransferase. Since the saccharide primer has a sugar residue as the starting point of glycan biosynthesis, saccharides are elongated into saccharide primers by the action of a wide variety of glycotransferases through interactions with cells. To date, several sugar derivatives that can serve as substrates for glycosyltransferase have been reported. It has been found, for example, that p-nitrophenyl β-D-xyloside serves as a substrate for glycosyltransferase1, and that 4-methyl umbelliferyl β-D-xyloside inhibits the synthesis of glycolipids and glycosaminoglycan (GAG) 2. A saccharide of mucin-type O-glycan is elongated to p-nitrophenyl α-D-GalNAc 3. It has also been reported that peracetylated Xylβ1-6Gal-O-2-naphthol and peracetylated Galβ1-4GlcNAcβ-O-naphthalenemethanol serve as substrates for glycosylation reactions in cells 4,5, and that such compounds serve as inhibitors of the biosynthesis of endogenous glycans 4-7. Sugar derivatives and products are considered to inhibit glycan biosynthesis when remaining in cells.
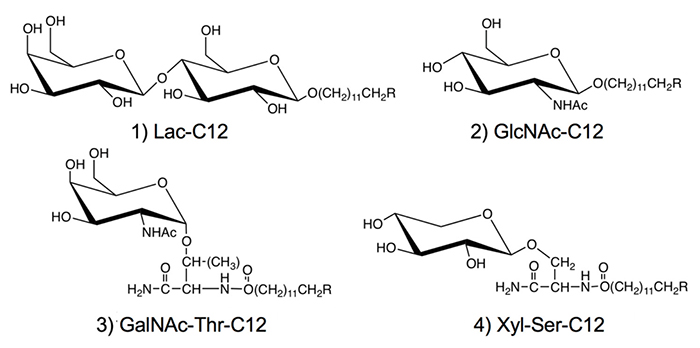
We used hydrocarbon chains such as dodecyl groups in the aglycone moiety of the saccharide primer (Fig. 1). With the optimized hydrocarbon chain length, the saccharide primer was incorporated in the cells, after which the saccharide primer and glycosylated products were not accumulated in the cells, but mostly secreted outside the cells 8. Therefore, the activity as an inhibitor of glycotransferase was low, and the glycosylation reaction was observed preferentially. Saccharide primers having an alkyl group are advantageous from the viewpoint of glycan synthesis, since glycosylated products can be recovered without cell homogenization because they are secreted extracellularly. To date, lactose (Lac) 8-11, N-acetylglucosamine (GlcNAc)12, N-acetylgalactosamine (GalNAc), and xylose (Xyl) 13-16 have been used as sugar residues. Since GalNAc and Xyl serve as precursors for the biosynthesis of O-glycan, glycoamino acids with Ser or Thr residue joined to sugar residue are used as glycotransferase recognition sites.
Since the saccharide primers shown in Figure 1 are characterized by extracellular secretion of products, we have been working to build a glycan library. In the synthesis for creating a glycan library, organic synthesis and enzymatic synthesis have commonly been performed. In contrast, in the saccharide primer method, a glycan library is created by making the best use of cells as a glycan synthesis factory. Hence, the method allowing cells to create a glycan library may be said to be the third method, following organic synthesis and enzymatic synthesis.
Our experiments on the generation of oligosaccharides in cells using the saccharide primer method follow the procedures illustrated below (Fig. 2). For example, cells are incubated in a 10-cm diameter culture dish, and when a subconfluent state is reached, the incubation medium is replaced with a serum-free medium containing 50 µM of a saccharide primer. After culturing for 2 days, the glycosylated product secreted in the culture medium is recovered. Since the product can be recovered without destroying the cells, the saccharide primer may be repeatedly added to recover the product. The recovered medium components are desalinized using a reversed-phase column, and the product is extracted in a solid form. It is also possible to separate acidic and neutral products by changing the composition of the eluting solvent. The chemical structure of the resulting products is analyzed using a mass spectrometer. Since the solid-extracted glycosylated product is a mixture of oligosaccharides, its structure is analyzed using LC-MS. For the liquid chromatography (LC), the products are separated using a silica column, etc. In the mass spectrometry (MS), MS2 spectra are measured using a mass spectrometer of the electrospray ionization / ion trap (ESI-IT) type, quadrupole time-of-flight (Q-TOF) type, or the like, followed by sequence analysis.
Sugar chain analysis by LC-MS is advantageous as it enables the separation of structural isomers by LC. In the case of NeuAc-Gal, for example, it is necessary to distinguish between the α2-3 and α2-6 bonds. LC retention time differs between the two cases, NeuAcα2-3 Gal having a shorter retention time. In addition, the two binding modes can be distinguished on the basis of the fragment ions detected by MS2 measurement. In the case of the NeuAcα2-3Galβ1-4GlcNAc-C12 bond, for example, MS2 spectrum peaks were detected at m/z of 823.4 and 550.2, whereas for the NeuAcα2-6Galβ1-4GlcNAc-C12 bond, these peaks were weakly detected, but m/z 335.9 was also detected, intensely. In the case of the Gal-GlcNAc bond, the binding modes of β1-3 and β1-4 must be distinguished. LC retention time is short for Galβ1-3GlcNAc and long for Galβ1-4GlcNAc. Furthermore, the MS2 spectra of Galβ1-4GlcNAc and Galβ1-3GlcNAc differ. As stated above, structural isomers, which have the same molecular weight, can be conveniently distinguished by analyzing LC elution times and MS2 spectra in LC-MS measurement. Although LC-MS is a useful method for identifying saccharide structures, it is sometimes difficult to determine the binding mode for oligosaccharides of low abundance, because of limitations of analyzer sensitivity.

In addition to distinguishing sugar binding modes, distinguishing monosaccharide structural isomers is also a difficult part of the structural analysis of glycans. In the case of hexose (Hex), for example, glucose (Glc), galactose (Gal), and mannose (Man) often need to be distinguished. For N-acetylhexosamine (HexNAc), N-acetylglucosamine (GlcNAc) and N-acetylgalactosamine (GalNAc) often need to be distinguished. Structural isomers of monosaccharides have conventionally been differentially identified using gas chromatography. If the differential identification of structural isomers of monosaccharides could be carried out using a mass spectrometer, it would be possible to improve the accuracy of the sequence analysis of glycans. Hence, we attempted to differentially identify structural isomers by ESI-IT-MS 17.
To distinguish structural isomers of monosaccharides, MS operating conditions were examined. The solvent effect was significant, and use of a sugar solution containing ammonium acetate was useful. Using a selected appropriate precursor ion when measuring MS2 and MS3 spectra in the positive ion mode, we also found that the fragment ion intensity ratio differed depending on the variety of monosaccharide (Table 1)17. The difference between hexose Glc and Gal, for example, was distinguishable by the difference in the intensity ratio of m/z 163 and m/z 180 in MS2 spectrum with a precursor ion of m/z 198. Gal and Man were differentially identifiable by measuring their MS3 spectra. Although the difference between GlcNAc and GalNAc was indistinguishable by examination of their MS2 spectra, they became distinguishable by measuring the intensity ratio of three fragment ions (m/z=126, 168, and 186) by MS3 spectrometry. In the negative ion mode, the isotopes were indistinguishable. As proposed above, the ESI-IT mass spectrometer enabled structural isomers to be analyzed under selected operating conditions.
The differential identification of monosaccharide structural isomers was also applicable to the analysis using oligosaccharides. As an example, we performed measurements with Gal-GlcNAc derivatives. MS2 spectrometry detected a fragment ion having a molecular weight corresponding to HexNAc. With this ion as the precursor ion, MS3 spectrometry detected a fragment ion at m/z 204. Next, the MS4 spectrum was measured with m/z 204 as the precursor ion. A fragment ion characteristic of GlcNAc was detected as shown in Table 1. To distinguish between GalNAc and GlcNAc in oligosaccharide sequence analysis, enzyme treatment is usually needed. As this experimental complexity can be avoided, ion trap type mass analyzers capable of measuring MSn spectra are considered to be useful in structural analysis of glycans. However, MS4 spectrometry requires sufficient sample concentration to obtain signal intensity, and analyzer sensitivity is also influential.
| Type of sugar | MS2 spectrum | MS3 spectrum | ||
| Precursor ion | Fragment ion intensity | Precursor ion | Fragment ion intensity | |
| Glc | m/z 198 [M+NH4]+ | m/z 163 < m/z 180 | m/z 163 | m/z 127 < m/z 145 |
| Gal | m/z 163 < m/z 180 | m/z 127 ≈ m/z 145 | ||
| Man | m/z 127 < m/z 145 | |||
| GlcNAc | m/z 222 [M+H]+ | m/z 186 < m/z 204 | m/z 204 | m/z 126 ≤ m/z 186 m/z 126 > m/z 168 |
| GalNAc | m/z 126 < m/z 186 m/z 126 < m/z 168 |
|||
The saccharide primer shown in Figure 1 is a precursor in various glycan biosynthetic pathways. The cells used express different glycan biosynthetic pathways depending on cell line. Therefore, a wide variety of glycans can be synthesized by combining a saccharide primer and cells. We call this a “biocombinatorial synthesis” method. We have so far applied saccharide primers to more than 50 types of cells, and have detected more than 200 different oligosaccharides. Major oligosaccharides obtained using saccharide primers Lac-C12, GlcNAc-C12, and Xyl-Ser-C12 are shown in Figures 3 to 5, respectively.
Lac-C12 is a saccharide primer for the precursor of the biosynthetic pathway for glycolipids. Therefore, oligosaccharides as found in major glycolipids, i.e., gangliosides, globo-series glycolipids, and neolacto-series glycolipids are elongated into the saccharide primer (Fig. 3). In B16 melanoma cells, a GM3 type product alone was observed as the only acidic sugar. In some other cells, disialoganglioside and trisialoganglioside were observed. Thus, oligosaccharides corresponding to glycolipids expressed in cells were also detectable using the saccharide primer method. In ganglio-series saccharides, a structure bound with glycoyl type sialic acid was also observed.
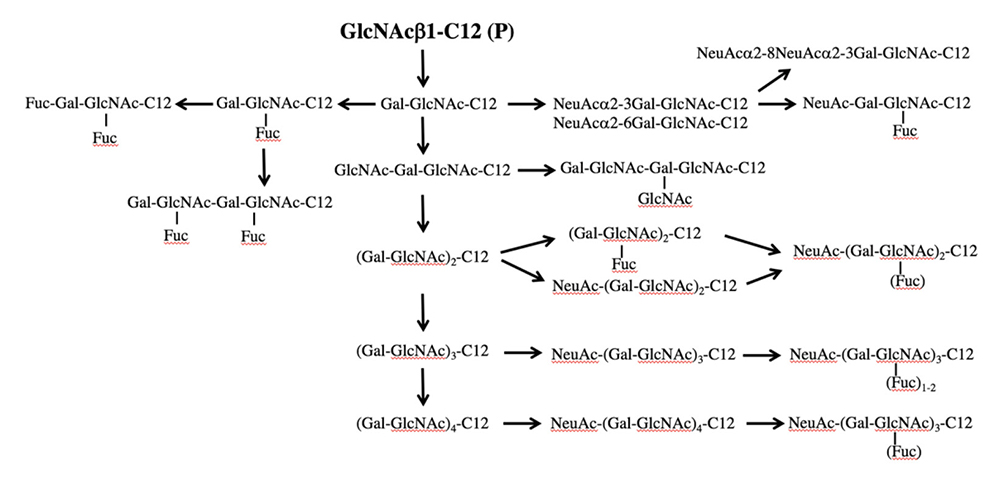
GlcNAc-C12 is a precursor of the biosynthetic pathway of lacto/neolacto-series glycans, such as lactosamine and sialyl Lewis antigen. When GlcNAc-C12 interacted with HL60 12 and other cell lines, Lewis X, sialyl Lewis X, polylactosamine, and sialylated polylactosamine were obtained (Fig. 4). The mode of binding of sialic acid to lactosamine included both α2-3 and α2-6, which were detected separately by LC-MS. Galβ1-4GlcNAc-C12 (LacNAc-C12) was also used as a saccharide primer to obtain neolacto-series glycans. As a result, a product similar to GlcNAc-C12 was obtained even with LacNAc-C12, but with only half the yield. Although there is a room for improvement, since the hydrophilicity-hydrophobicity balance of the saccharide primer seems to have an influence, we consider that monosaccharide-type GlcNAc-C12 is suitable as a saccharide primer for obtaining neolacto-series glycans.
The saccharide primer Xyl-Ser-C12 is a precursor of the biosynthesis of proteoglycan-type glycans. In the biosynthesis of proteoglycan, Gal-Gal is added to the Xyl residue, beyond which HexA and HexNAc are repeatedly elongated to form the (HexA-HexNAc)n-Gal-Gal-Xyl structure. When CHO cells were interacted with Xyl-Ser-C12, Gal-Gal-Xyl-Ser-C12 was produced, and then an elongation product of HexNAc and HexA, i.e., (HexNAc-HexA)n-Gal-Gal-Xyl-Ser-C12, was detected; a proteoglycan-type glycan was obtained (Fig. 5). In CHO cells, elongation of 10 sugars was found in the saccharide primer at up to n=4, and 14 products were obtained 13. To confirm whether Xyl-Ser is advantageous in the biosynthesis of proteoglycan-type glycans, we conducted a comparative experiment with Xyl-pNP. When CHO cells were interacted with Xyl-pNP, only 5 saccharide elongation products were detected under the same conditions as with Xyl-Ser-C12. The presence of the Ser residue in Xyl-Ser-C12 was useful from the viewpoint of glycan synthesis. Hence, we synthesized Xyl-Thr-C12 replaced with Thr and found almost no elongation of saccharides; it was thus suggested that the presence of Ser as the xylose-binding amino acid residue was important to the recognition of glycotransferase 15. Furthermore, in view of the consensus motif of Xyl-binding amino acids, we designed Gly-(Xyl)Ser-C12, which has Gly-Ser in its amino acid residue. As a result, sialyloligosaccharides such as NeuAc-Gal-Xyl-Ser-C12 were detected in the glycosylated products from Xyl-Ser-C12, whereas the sialylation reaction was suppressed in the case of Gly-(Xyl)Ser-C12 15. While NeuAc modifications are considered to inhibit the elongation of proteoglycan-type glycans, it was demonstrated that the elongation of proteoglycan-type glycans occurred preferentially as sialic acid transfers were suppressed by Gly-Ser residues.
In normal human dermal fibroblast NHDF cells, the sialyloligosaccharide NeuAcα2-3Gal-Xyl-Ser-C12 was the major product, with phosphorylated NeuAcα2-3Gal-Xyl2P-Ser-C12 detected as the second-most abundant product 15. In addition, NeuAcα2-6Gal-Gal-Xyl-Ser-C12, a structure estimated to be an intermediate for GAG chain synthesis, was also detected. In the GAG synthetic pathway, phosphorylated HexNAc-HexA-Hex-Hex-Xyl2P-Ser-C12 was most abundantly detected. In addition, a product with HexNAc-HexA repeats, i.e., (HexNAc-HexA)2-Hex-Hex-Xyl2P-Ser-C12, was produced, and this product is hydrolyzed by C-ABC but not by heparitinase, which indicated the elongation of the GAG chain of the chondroitin sulfate type (GalNAc-GlcA)n-Gal-Gal-Xyl2P-Ser-C12. On the other hand, a product with dephosphorylated Xyl, i.e., (HexNAc-HexA)2-Hex-Hex-Xyl-Ser-C12, was hydrolyzed by both C-ABC and heparitinase. Hence, it was conjectured that the GAG chains of the dephosphorylated chondroitin sulfate type (GalNAc-GlcA)n-Gal-Gal-Xyl-Ser-C12 and the heparan sulfate type (GlcNAc-GlcA)n-Gal-Gal-Xyl-Ser-C12 were elongated. Judging from LC-MS peak intensity, the major product was identified as the chondroitin sulfate type. This suggested that chondroitin sulfate elongated with Xyl phosphorylation, and that Xyl dephosphorylation was needed for the elongation of the heparan sulfate type. This demonstrated the role of xylose phosphorylation in the GAG chain elongation reaction; detection of intermediates for the biosynthetic pathway for GAG chains is considered to be effective in analyzing the mechanism of the biosynthesis.
Furthermore, saccharide primers having GalNAc, i.e., GalNAc-Thr-C12 and GalNAc-Ser-C12, were synthesized to acquire mucin-type glycans. When they were interacted with more than one cell line, products classified as core 1 and core 2, including sialyl Tn antigen, T antigen, sialyl T antigen, and fucosyl T antigen, were mainly detected in many cells. These saccharide primers are expected to be efficiently used to build a mucin-type glycan library.


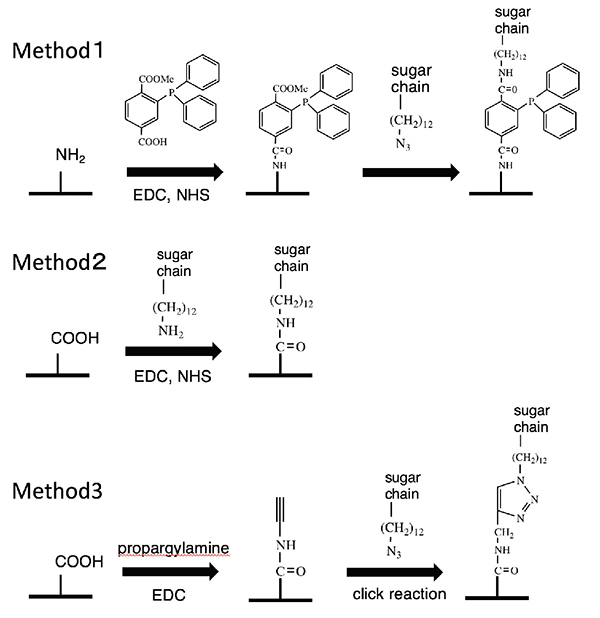
Enabling the synthesis of a wide variety of glycans, this saccharide primer method makes it possible to build a glycan library. To confirm the glycan typing, the resulting glycan must be immobilized on a sensor or the like. To this end, we have developed an azidated saccharide primer 18. The azido group introduced into the aglycone end does not influence saccharide elongation reactions. This allows cells to synthesize a glycan library having the azido group. The azidated glycan library can be chemically immobilized on solid surfaces. To achieve this, we carried out 1) a Schtaudinger reaction 19, 2) reduction of the azido group to the amino group followed by condensation reaction 19, and 3) a click reaction, as shown in Figure 6.
In AIDS virus (HIV) infection, it is known that binding of envelope proteins (gp120, gp160) and glycolipids induces the fusion of HIV and biomembranes 20. In particular, gp120 has been reported to have affinity for GM3, GalCer, or Gb3Cer 21-23. Therefore, we synthesized Gal-Gal-Glc-C12-N3 having Gb3 or GM3-type NeuAc-Gal-Glc-C12-N3 through interaction of Lac-C12-N3 with mouse B16 melanoma cells. This compound was immobilized using a condensation reaction with the carboxy group of the sensor substrate by reducing its azido group to an amino group. The interaction between the immobilized oligosaccharide and gp120 or gp160 was quantitatively evaluated using a surface plasmon resonance apparatus. As a result, gp160 showed affinity for Gb3 and Lac rather than for GM3-type, and gp120 showed equally low affinity for all the oligosaccharides examined 24.
Recently, we have succeeded in detecting influenza viruses by immobilizing a saccharide library obtained using the saccharide primer method on microparticle surfaces by a click reaction. Hence, the saccharide primer method enables an azidated glycan library to be synthesized conveniently, and can be utilized to evaluate carbohydrate recognition by the best use of the library.
As stated above, we have been working on a saccharide primer method with the aim of building a glycan library. Additionally, the saccharide primer method enables the detection of glycosylated products via cellular glycan biosynthetic pathway with a small number of cells, and is therefore applicable to analysis of the expression of glycans related to cell functions. Some example applications used in the past are described below.
Neuroblastoma is a cancer that develops in nerve cells with high incidence in children aged 5 years and younger. Neuroblastoma cells exhibit higher expression of GD2 than normal nerve cells 25. In addition, a relation is known to exist between ganglioside expression and prognosis; the prognosis was reported to be good with the expression of gangliosides of the ganglio-b series 26, and to be poor with low expression of gangliosides of the ganglio-b series 27.
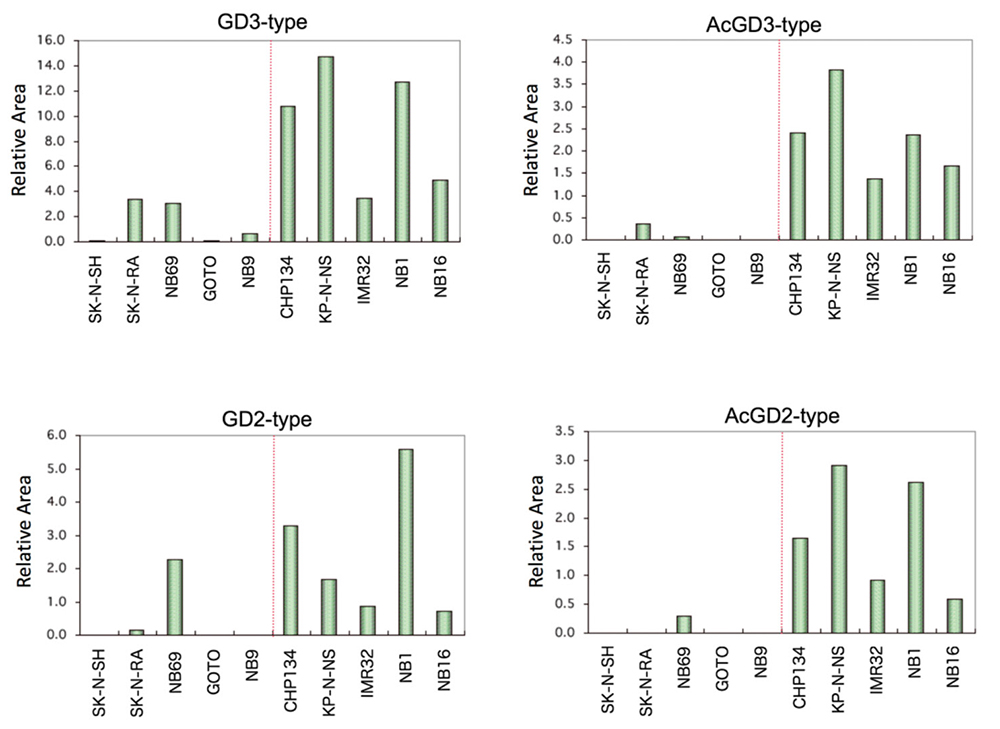
We induced interaction of the saccharide primer Lac-C12 in 10 neuroblastoma cell lines with various expression levels of nerve differentiation markers (Phox2a/b, TrkC, NCAM, neurofilament, etc.), and analyzed the chemical structures of the resulting saccharide elongation products. A total of 16 glycosylated products were detected, including the ganglioside series and globo series. In particular, the cell lines with high expression levels of the GD3 type, GD2 type, and their acetylated products showed high expression of nerve differentiation markers (Fig. 7). When endogenous glycolipids were analyzed using the same cell line, GD2 and acetylated GD2 were shown to correspond to the expression of nerve differentiation markers 28. In addition, analysis of glycogenes demonstrated high expression of the GD3 synthase gene ST8SIA1 in the cell lines with high expression of nerve differentiation markers. On the other hand, no association was found between the expression of the GalNAc-transferase gene B4GALNT1, Gal-transferase gene B3GALT4, or NeuAc-transferase gene ST3GAL1/4 and nerve differentiation markers. This suggests that the improved expression of ganglio-b-series glycans in neuroblastoma was influenced by the action of ST8SIA1 to improve the expression of GD3.
Human promyelocytic leukemia cell line HL-60 is capable of differentiating to monocytic or granulocytic cells. When treated with 12-O-acetyldecanoylphorbol-13-acetate (TPA), for example, HL-60 differentiates to monocytic cells. This is known to markedly increase GM3 and to decrease neolacto-series glycolipid nLc4Cer 29. Therefore, we induced interaction of Lac-C12 and GlcNAc-C12 in HL-60 cells before and after inducing their differentiation, and measured the changes in the resulting glycosylated products by LC-MS. The cells became adhesive 12 hours after addition of TPA and exhibited non-specific esterase activity at 72 hours, demonstrating induced differentiation. Around the time when cells exhibited adhesive behavior, the detection of GM3-type (NeuAc-Gal-Glc-C12) and Gb3(Gal-Gal-Glc-C12) improved in Lac-C12-derived products. In GlcNAc-C12-derived products, on the other hand, the synthesis of Gal-(Fuc)GlcNAc-C12 began to decrease. These changes in saccharide expression levels were correlated with the changes in the expression levels of genes such as the GM3 synthase gene, Gb3 synthase gene, and FUT4. After addition of TPA, the expression of the GM3 type first rose, and then the expression of Gb3 rose; when non-specific esterase activity was found, their expression levels peaked. As the GM3 type and Gb3 increased, Gal-(Fuc)GlcNAc-C12 began to decrease. Hence, the saccharide primer method was shown to enable convenient monitoring of changes in glycan biosynthesis following addition of the differentiation induction reagent.
Embryonic carcinoma cells (EC cells) retain their undifferentiated state in teratocarcinoma, a naturally developing tumor in the testis and ovary. Because of their similarly to early embryo cells, EC cells have been used as differentiation model cells in research on the early developmental process. F9 cells, a type of mouse ES cell, are known to exhibit characteristics similar to those of proximal endoderm when treated with retinoic acid (RA)29, and exhibit characteristics similar to those of distal endoderm when treated with RA and dibutyryl cyclic AMP (dcAMP) 30.
It is known that the SSEA 1/3 and the Forssman antigens are expressed in F9. When F9 is induced to differentiate with RA, the expression of SSEA1 31 and Forssman antigens 32 decrease, and SSEA3 33 and ganglioside 29,34 increase. We examined changes in the glycan biosynthetic pathway by interacting GlcNAc-C12 with F9 cells induced to differentiate by the action of RA and dcAMP 35. As a result, the amount of neolacto-series glycans from GlcNAc-C12 synthesized increased as a whole. NeuAc-Gal-GlcNAc-C12, in particular, increased markedly. Furthermore, expression of synthase genes for the glycans obtained, B4galt1, which is for the synthesis of Galβ1-4GlcNAc, and St3gal6, which is for the synthesis of NeuAcα2-3Gal, improved. At the same time, endogenous glycolipids were extracted and analyzed using a mass spectrometer; increased expression of paragloboside (nLc4Cer) and sialylparagloboside was found.
As a factor for cancer malignancy, the relationship between metastatic potential and glycans has been studied extensively 36-41. For example, the interaction between tumor cells and vascular endothelial cells is involved by sialyl Lex antigen and sulfated sialyl Lex antigen, and tissue infiltration is involved by gangliosides and sulfated glycolipids. Heparan sulfate is also known to be involved in tumor cell growth and metastasis. We attempted to identify glycans involved in the metastasis of mouse osteosarcoma cells. To this end, we induced the interaction of Xyl-Ser-C12 with FBJ-LL, a mouse osteosarcoma cell line of high metastatic potential, and FBJ-S1 cells of low metastatic potential, and analyzed glycan 15. A glycosylated product with Gal-Gal-Xyl-Ser-C12 bound by repeats of HexA and HexNAc was obtained from these osteosarcoma cells. Since this product was hydrolyzed by heparitinase, it was estimated to be GAG of the heparan sulfate type. Although a nearly identical product was obtained from FBJ-LL cells and FBJ-S1 cells, the amount of the product of the GAG type was significantly lower in FBJ-LL cells of high metastatic potential. When evaluated by RT-PCR and western blotting, the expression of GAG-chain extension enzymes (Ext1 and Ext2) was shown to be lower in FBJ-LL cells than in FBJ-S1 cells, demonstrating correspondence to the amount of glycosylated product detected. Hence, the Ext1 gene was knocked down in FBJ-S1 cells, which showed high expression of Ext1, using the siRNA method; cell migration improved. Furthermore, unexpectedly, increased expression of heparanase was also observed in the FBJ-S1 cells with the knocked down Ext1 gene. Thus, the expression of heparan sulfate was shown to be linked to its synthesis and hydrolysis. We were therefore able to assess the relationship between cell migration and glycans using the saccharide primer method.
We have been working on two approaches to the best use of the saccharide primer method: one is to generate a glycan library, and the other is to profile glycans expressed in cells. These two goals can be attained at one time by interacting a saccharide primer with cells, and analyzing the resulting glycosylated products by LC-MS. We are making the best use of the glycan library thus obtained to analyze carbohydrate recognition for proteins and viruses, to develop a database for glycan structural analysis by LC-MS, and for other purposes. We are also working to develop a database of glycans expressed in cells by profiling glycans in a wide variety of cells, and to apply the information to comparative analysis and correlation with cell functions, using bioinformatic techniques. By doing so, we have become able to identify glycans depending on cell type and function, and to make an efficient approach to the analysis of genes involved in the synthesis of such glycans. We hope that our saccharide primer method, as a novel tool, will contribute to the development of glycomics.