
Takashi Sato
Takashi Sato was born in Fukushima. He graduated from the University of Utsunomiya in 1994 and obtained his Ph.D. from the University of Tsukuba in 1999. He participated in the GlycoGene (GG) Project in 2001 and the Structual Glycomics (SG) Project in 2003 at the National Institute of Advanced Industrial Science and Technology (AIST). His recent research interests are protein specific glycosylation mechanisms and glycosyltransferases.

Masanori Gotoh
Masanori Gotoh graduated from the Faculty of Science and Technology of the Tokyo University of Science in 1985 and completed his master’s degree course at the same University in 1987. He received his Ph.D. from the University of Tsukuba in 2003. He worked at Toshiba Corporation and Pharmacia Biotech KK for six and eight years, respectively. He took part in the Construction of Libraries of Human Genes Participating in Glycosylation Project at the Research Center for Glycoscience of National Institute of the Advanced Industrial Science and Technology (AIST) in 2001. At AIST, he took part in the establishment of GlycoGene, Inc., in 2005 and holds the additional post of Chief Technology Officer.

Hisashi Narimatsu
Hisashi Narimatsu was born in Naruto city and graduated from the Keio University School of Medicine and obtained M.D. in 1974. He obtained his Ph.D. in the Department of Microbiology at the Keio University Postgraduate School of Medicine in 1979. He succeeded in cloning the β4Gal-T1 gene, the first glycosyltransferase gene cloned in the world in Dr. Pradman Qasba’s laboratory at the U.S. National Institute of Health in 1985. He returned to the Department of Microbiology of Keio University as assistant professor in 1986. In 1991 he became a professor at the Institute of Life Science, Soka University, after which he moved to the National Institute of Advanced Industrial Science and Technology (AIST) and founded the Research Center for Glycoscience (RCG) in 2002. He is currently leader of the Glycogene Function Team, Deputy Director of RCG and Professor of the Graduate School of Comprehensive Human Sciences, University of Tsukuba.
Glycosyltransferase genes can be classified into several families according to amino acid sequence homology. Each family has its own motif sequence, some of which have recently been identified as essential amino acids for binding to donor and acceptor substrates. Using the sequence of β1,4- galactosyltransferase (β4Gal-T) as a query, we searched for novel glycosyltransferases having a β1,4-linkage motif (β4GT motif, WGXEDD/V/W) in databases, and found seven novel sequences. Five of them were determined to be the glycosyltransferase genes involved in the synthesis of chondroitin sulfate, and the other two genes were determined to be β1,4-N-acetylgalactosaminyltransferases. At the present time, 15 members of the β4GT family with a β1,4-linkage motif have been identified (Fig. 1). This article reviews the glycosyltransferase family with the β1,4-linkage motif, and recent findings regarding the substrate specificities of seven β1,4-galactosyltransferases (β4Gal-T) and two β1,4-N-acetylgalactosaminyltransferases (β4GalNAc-T), other than the chondroitin sulfate synthase family (see under separate heading).
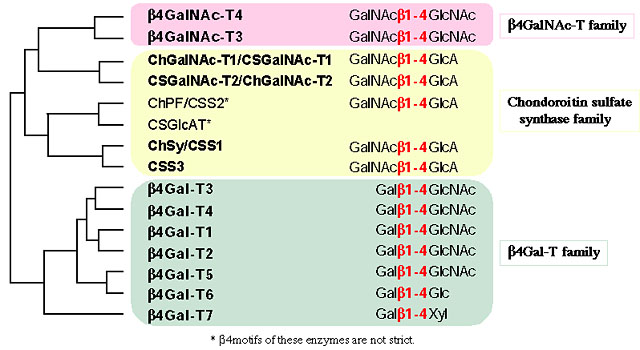

Fig. 1 A phylogenetic tree and substrate specificity of the β4GT family
Bioinformatics has enabled researchers to effectively find novel candidate genes from enormous human genome sequences and EST databases. First of all, we searched motif sequences conserved in the β4GT family with bioinformatics technology. The motif WGXEDD/V/W was found to be conserved in all members reported. In addition to this motif, some typical features to the glycosyltransferases, such as the DXD motif, which is known to be a binding motif to divalent cations, N-terminal hydrophobic transmembrane region, were employed as parameters for bioinformatics search. Using these parameters, we searched databases and found partial sequences of novel glycosyltransferase genes. Based on the partial sequences, we determined the entire amino acid sequences and performed expression of recombinant enzymes. We found that all enzymes transferred sugars via a β1,4-linkage by analyzing the substrate specificities of each enzyme. Fig. 1 briefly shows the substrate specificities and results of phylogenetic analysis of novel glycosyltransferases genes and β1,4Gal-T genes. As shown in Fig. 2, an entire family conserves the DXD motif and the WGXEDD/V/W region (β4GT motif). Ramakrishnan et al. performed X-ray crystallographic analysis of β1,4Gal-T1 and reported that this WGXEDD/V/W motif might be involved in the binding to a substrate 1.
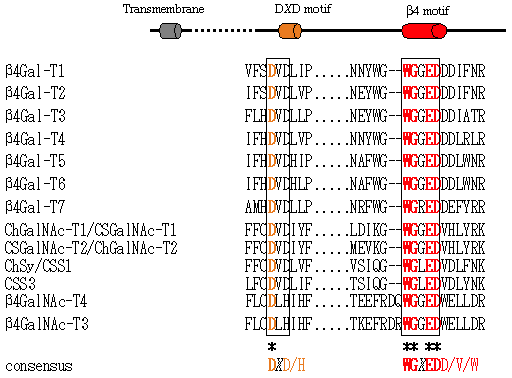
Fig. 2 Motif sequences conserved in the β4GT family
Seven members of β4Gal-T gene family have been reported. β4Gal-T1 is the first glycosyltransferase gene cloned 2 and is one of the enzymes that have been analyzed most extensively. β4Gal-T1 has the enzyme activity of transferring Gal from a donor substrate UDP-Gal to an acceptor substrate GlcNAcβ- and has the activity of synthesizing lactose (Galβ1-4Glc) with lactalbumin. Glycosyltransferase activities towards glycolipids and N- or O-linked glycan of glycoproteins have been reported. Although β4Gal-T2, -T3, -T4, and –T5 synthesize Galβ1-4GlcNAc- using UDP-Gal, like β4Gal-T1, in vitro analyses have shown other substrate specificities. β4Gal-T2 has substrate specificity very similar to that of β4Gal-T1, whereas β4Gal-T3 uses Lc3Cer as an acceptor substrate. β4Gal-T4 is thought to be keratan sulfate synthase because it uses GlcNAc-6-sulfate, which is a component of keratan sulfate, as the substrate. β4Gal-T6 uses Glc-Cer as the acceptor substrate to synthesize Lac-Cer. β4Gal-T7 uses Xyl-Ser as the acceptor substrate to synthesize the Galβ1-4Xyl-Ser present in the common structure at the linkage region of chondroitin sulfate and heparan sulfate. We briefly described the substrate specificities of the β4Gal-T family. For more details, the reader is referred to other reviews 3-5.
β4GalNAc-T3 and β4GalNAc-T4 belong to the β4GalNAc-T gene family. These enzymes are type II membrane proteins that have a transmembrane region at the N-terminus and no homology to other glycosyltransferases except for a β1,4-linkage motif. These genes were found to encode large ORFs for 999 and 1039 amino acids, but did not have two glycosyltransferase domains as observed in chondroitin sulfate synthase. The predicted catalytic domain of the glycosyltransferases would be in the C-terminal region, and it contained a β1,4-linkage motif and a DLH motif which appeared to have a function similar to that of DXD (Fig. 2). However, the N-terminal region had no homology to any known amino acid sequences.
Analysis of the substrate specificities of these enzymes showed that they transfers GalNAc from the donor substrate UDP-GalNAc to the acceptor substrate GlcNAcβ1- via a β1,4-linkage to synthesize the GalNAcβ1,4GlcNAcβ1-(LacdiNAc-) structure. The LacdiNAc structure is a special structure that has been specifically found on N-glycans of glycoprotein hormones such as luteinizing hormone (LH) and glycodelin (Gd). The sulfation, fucosylation and sialylation occurr toward LacdiNAc (Fig. 3), and the sulfated LacdiNAc plays an important role in the clearance of glycoprotein hormones. These enzymes were found to use the biantennary structures of N-glycan as an acceptor in vitro (Table 1). They differed greatly in expression tissues. β4GalNAc-T3 was expressed in the stomach and colon, whereas β4GalNAc-T4 was highly expressed in the brain and ovary, suggesting that the LacdiNAc structure found on LH and Gd may be synthesized by β4GalNAc-T4 6,7.
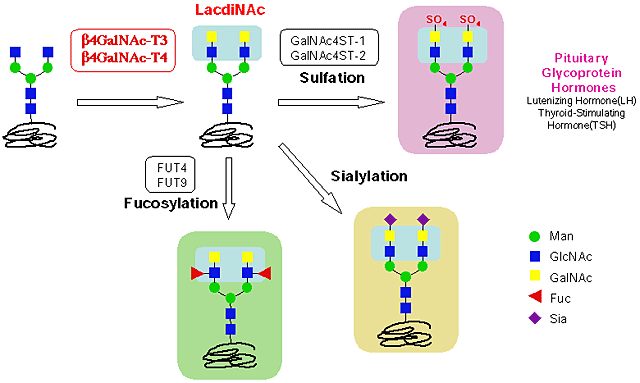
Fig. 3 LacdiNAc Synthesis and its Modification on N-glycans
Table 1 Substrate specificity of β4GalNAc-Ts toward N-glycans

We have briefly described a set of glycosyltransferases with a β4GT motif. The human genome does not contain sequences that encode proteins with a β1,4-linkage motif. Other glycosyltransferases such as β4GalNAc-T1 and β4GalNAc-T2, synthesizing a β1,4-linkage without definite β4GT motif have been found 8,9. Since many β1,4-linked glycan structures exist in the human, more detailed analysis is needed for the substrate specificities of known glycosyltransferases and of unknown glycosyltransferases that may be discovered in the future.