|
Phage Display Method:
Analysis of the interaction between ganglioside and peptide
|
|
|
 |
In 1993, G. P. Smith et al. developed a method for efficiently selecting peptide sequences that can bind to the target molecule from the peptide library. This method is called the phage display method, because the peptide library is displayed on the outer shell protein of the filamentous phage (1). The peptide sequences expressed in the outer shell protein can be easily identified by sequence analysis of the phage DNA (Figure). The library is presented as proteins such as antibody or enzyme, linear or cyclic peptides, and so on. The numbers of presented molecules are 3-5 copies for the pIII protein (the minor coat protein), and 2700 copies for pVIII (the major coat protein). Since the size of the library is about 108-109, the number of amino acid residues is 6-7 in the calculation including all combinations (206=6.4x107). Therefore, a random library of 15 amino acids can not contain all of the sequences. Proteins such as antibodies or receptors, nucleic acids, glycoconjugates, low molecular weight compounds, cells, organs, plastics, etc. are used for the target molecule (2). The phage library technique is used to obtain target-specific molecules, and to know the epitope of the peptide sequence that is related to carbohydrate recognition. Phage display libraries are commercially available at reasonable prices. |
|
|
 |
|
|
Using the phage display method, it is possible to analyze the interaction between sugar chains in the ganglioside and amino acids in the protein. By performing the selection of the carbohydrate-specific peptides from random peptide library, it is possible to specify the amino acid that is related to the carbohydrate recognition. Moreover, the amino acid sequence obtained from the selection may assist in understanding carbohydrate-protein interaction, if there is homology between the selected peptide and natural protein, and any regularity of epitope.
The immobilization of the target molecule affects the success of the selection. For example, to obtain the peptides that bind to the sugar chain of a glycolipid, it is important to use a highly ordered membrane, such as a biomembrane. In fact, a glycolipid-specific peptide was not selected when the selection was carried out using the disordered lipid membrane. On the other hand, specific peptide sequences were successfully selected by performing biopanning with the ordered glycolipid monolayer.
For example, biopanning for ganglioside GM1 (Galß1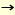 3 GalNAcß1 3 GalNAcß1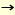 4 (NeuAca2 4 (NeuAca2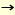 3) Galß1 3) Galß1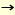 4Glcß1 4Glcß1 1'Cer) was performed using phage library, this presented a random 15 amino acid residues (Figure). As a result, only three kinds of peptides were selected (3). Though there was no homology between the natural sugar-binding protein and the selected peptide sequences, the common motif f/wRxL-Px-Fxx-R-xP was found in two of the three sequences. The peptides obtained inhibited the binding of the cholera toxin B subunit for GM1 at 1.0µM of IC50 (3). It is believed that roles of amino acids obtained R, F (or L), and P were correlated to the interaction with sialic acid, the stacking with galactose, and the bending of the peptide, respectively. Furthermore, it is possible to carry out the selection of peptides for other glycolipids besides GM1, and experiments to deduce the general law of the sequences necessary for the carbohydrate recognition are now in progress. 1'Cer) was performed using phage library, this presented a random 15 amino acid residues (Figure). As a result, only three kinds of peptides were selected (3). Though there was no homology between the natural sugar-binding protein and the selected peptide sequences, the common motif f/wRxL-Px-Fxx-R-xP was found in two of the three sequences. The peptides obtained inhibited the binding of the cholera toxin B subunit for GM1 at 1.0µM of IC50 (3). It is believed that roles of amino acids obtained R, F (or L), and P were correlated to the interaction with sialic acid, the stacking with galactose, and the bending of the peptide, respectively. Furthermore, it is possible to carry out the selection of peptides for other glycolipids besides GM1, and experiments to deduce the general law of the sequences necessary for the carbohydrate recognition are now in progress.
Selection technology by this phage display method can also be used to obtain sugar-replica peptides (4). The sugar-replica peptide may be utilized as inhibitors for the sugar-binding proteins and the cells expressing the proteins. The phage display method will become an ever more useful technique as carbohydrate research advances.
|
|
|
|
Teruhiko Matsubara
(Department of Chemical Science and Technology, The University of Tokushima)
Toshinori Sato
(Department of Applied Chemistry, Keio University)
|
|
|
|
|
|
| References |
(1) |
Smith GP, Scott JK, Methods Enzymol. 217, 228-257, 1993 |
|
(2) |
Smith GP, Petrenko VA, Chem. Rev. 97, 391-410, 1997 |
|
(3) |
Matsubara T, Ishikawa D, Taki T, Okahata Y, Sato T, FEBS Lett. 456, 253-256, 1999
|
|
(4) |
Ishikawa D, Kikkawa H, Ogino K, Hirabayashi Y, Oku N, Taki T, FEBS Lett. 441, 20-24, 1998
|
|
|
|
|
|
| Sep.15, 2001 |
|
|
|
|
|
|
|



