|
| References |
(1) |
Habuchi O, Matsui Y, Kotoya Y, Aoyama Y, Yasuda Y, Noda M, Purification of chondroitin 6-sulfotransferase secreted from cultured chick embryo chondrocytes. J. Biol. Chem. 268, 21968-21974, 1993 |
|
(2) |
Fukuta M, Uchimura K, Nakashima K, Kato M, Kimata K, Shinomura T, Habuchi O, Molecular cloning and expression of chick chondrocyte chondroitin 6-sulfotransferase. J. Biol. Chem. 270, 18575-18580, 1995 |
|
(3) |
Habuchi O, Hirahara Y, Uchimura K, Fukuta M, Enzymatic sulfation of galactose residue of keratan sulfate by chondroitin 6-sulfotransferase. Glycobiology 6, 51-57, 1996 |
|
(4) |
Fukuta M, Kobayashi Y, Uchimura K, Kimata K, Habuchi O, Molecular cloning and expression of human chondroitin 6-sulfotransferase. Biochim. Biophys. Acta 1399, 57-61, 1998 |
|
(5) |
Yamada T, Ohtake S, Sato M, Habuchi O, Chondroitin 4-sulphotransferase-1 and chondroitin 6-sulphotransferase-1 are affected differently by uronic acid residues neighbouring the acceptor GalNAc residues. Biochem. J. 384, 567-575, 2004 |
|
(6) |
Kitagawa H, Fujita M, Ito N, Sugahara K. Molecular cloning and expression of a novel chondroitin 6-O-sulfotransferase. J. Biol. Chem. 275, 21075-21080, 2000 |
|
(7) |
Fukuta M, Inazawa J, Torii T, Tsuzuki K, Shimada E Habuchi O, Molecular cloning and characterization of human keratan sulfate Gal-6-sulfotransferase. J. Biol. Chem. 275, 21075-21080, 2000 |
|
(8) |
Uchimura K, Muramatsu H, Kadomatsu K, Fan Q-W, Kurosawa N, Mitsuoka C, Kannagi R, Habuchi O, Muramatsu T, Molecular cloning and characterization of an N-acetylglucosamine-6-O-sulfotransferase. J. Biol. Chem. 273, 22577-22583, 1998 |
|
(9) |
Uchimura K, Muramatsu H, Kaname T, Ogawa H, Yamakawa T, Fan Q-W, Mitsuoka C, Kannagi R, Habuchi O, Yokoyama I, Yamamura K, Ozaki T, Nakagawara A, Kadomatsu K, Muramatsu T, Human N-acetylglucosamine-6-O-sulfotransferase involved in the biosynthesis of 6-sulfo sialyl Lewis X: molecular cloning, chromosomal mapping, and expression in various organs and tumor cells. J. Biochem. 124, 670-678, 1998 |
|
(10) |
Akama TO, Nishida K, Nakayama J, Watanabe H, Ozaki K, Nakamura T, Dota A, Kawasaki S, Inoue Y, Maeda N, Yamamoto S, Fujiwara T, Thonar EJ, Shimomura Y, Kinoshita S, Tanigami A, Fukuda MN, Macular corneal dystrophy type I and type II are caused by distinct mutations in a new sulphotransferase gene. Nat. Genet. 26, 237-41, 2000 |
|
(11) |
Akama TO, Nakayama J, Nishida K, Hiraoka N, Suzuki M, McAuliffe J, Hindsgaul O, Fukuda M, Fukuda MN, Human corneal GlcNac 6-O-sulfotransferase and mouse intestinal GlcNac 6-O-sulfotransferase both produce keratan sulfate. J. Biol. Chem. 276, 16271-16278, 2001 |
|
(12) |
Yamauchi S, Hirahara Y, Usui H, Takeda Y, Hoshino M, Fukuta M, Kimura JH, Habuchi O, , J. Biol. Chem. 274, 2456-2463, 1999 |
|
(13) |
Yamauchi S, Mita S, Matsubara T, Fukuta M, Habuchi H, Kimata K, Habuchi O, Purification and characterization of chondroitin 4-sulfotransferase from the culture medium of a rat chondrosarcoma cell line. J. Biol. Chem. 275, 8975-8981, 2000 |
|
(14) |
Okuda T, Mita S, Yamauchi S, Matsubara T, Yagi F, Yamamori D, Fukuta M, Kuroiwa A, Matsuda Y, Habuchi O, Molecular cloning, expression, and chromosomal mapping of human chondroitin 4-sulfotransferase, whose expression pattern in human tissues is different from that of chondroitin 6-sulfotransferase. J. Biochem. 128, 763-770, 2000 |
|
(15) |
Hiraoka N, Nakagawa H, Ong E, Akama TO, Fukuda MN, Fukuda M, Molecular cloning and expression of two distinct human chondroitin 4-O-sulfotransferases that belong to the HNK-1 sulfotransferase gene family. J. Biol. Chem. 275, 20188-20196, 2000 |
|
(16) |
Mikami T, Mizumoto S, Kago N, Kitagawa H, Sugahara K, Specificities of three distinct human chondroitin/dermatan N-acetylgalactosamine 4-O-sulfotransferases demonstrated using partially desulfated dermatan sulfate as an acceptor: implication of differential roles in dermatan sulfate biosynthesis. J. Biol. Chem. 278, 36115-36127, 2003 |
|
(17) |
Kang HG, Evers MR., Xia G, Baenziger JU, Schachner M, Molecular cloning and characterization of chondroitin-4-O-sulfotransferase-3. A novel member of the HNK-1 family of sulfotransferases. J. Biol. Chem. 277, 34766-34772, 2002 |
|
(18) |
Evers MR, Xia G, Kang HG, Schachner M, Baenziger JU, J. Biol. Chem. 275, 34728-34736, 2000 |
|
(19) |
Ito Y, Habuchi O, Purification and characterization of N-acetylgalactosamine 4-sulfate 6-O-sulfotransferase from the squid cartilage. J. Biol. Chem. 275, 34728-34736, 2000 |
|
(20) |
Yamaguchi T, Ohtake S, Kimata K, Habuchi O, submitted, 2007 |
|
(21) |
Ohtake S, Ito Y, Fukuta M, Habuchi O, Human N-acetylgalactosamine 4-sulfate 6-O-sulfotransferase cDNA is related to human B cell recombination activating gene-associated gene. J. Biol. Chem. 276, 43894-43900, 2001 |
|
(22) |
Ohtake S, Kimata K, Habuchi O, A unique nonreducing terminal modification of chondroitin sulfate by N-acetylgalactosamine 4-sulfate 6-o-sulfotransferase. J. Biol. Chem. 278, 38443-38452, 2003 |
|
(23) |
Brandan E, Hirschberg CB, Purification of rat liver N-heparan-sulfate sulfotransferase. J. Biol. Chem. 263, 2417-2422, 1988 |
|
(24) |
Hashimoto Y, Orellana A, Gil G, Hirschberg CB, Molecular cloning and expression of rat liver N-heparan sulfate sulfotransferase. J. Biol. Chem. 267, 15744-15750, 1992 |
|
(25) |
Dixon J, Loftus SK, Gladwin AJ, Scambler PJ, Wasmuth JJ, Dixon MJ, Cloning of the human heparan sulfate-N-deacetylase/N-sulfotransferase gene from the Treacher Collins syndrome candidate region at 5q32-q33.1. Genomics 26, 239-244, 1995 |
|
(26) |
Humphries DE, Sullivan BM, Aleixo MD, Stow JL, Localization of human heparan glucosaminyl N-deacetylase/N-sulphotransferase to the trans-Golgi network. . Biochem. J. 325, 351-357, 1997 |
|
(27) |
Pettersson I, Kusche M, Unger E, Wlad H, Nylund L, Lindahl U, Kjell n L. Biosynthesis of heparin. Purification of a 110-kDa mouse mastocytoma protein required for both glucosaminyl N-deacetylation and N-sulfation. J. Biol. Chem. 266, 8044-8049, 1991 n L. Biosynthesis of heparin. Purification of a 110-kDa mouse mastocytoma protein required for both glucosaminyl N-deacetylation and N-sulfation. J. Biol. Chem. 266, 8044-8049, 1991 |
|
(28) |
Orellana A, Hirschberg CB, Wei Z, Swiedler SJ, Ishihara M, Molecular cloning and expression of a glycosaminoglycan N-acetylglucosaminyl N-deacetylase/N-sulfotransferase from a heparin-producing cell line. J. Biol. Chem. 269, 2270-2276, 1994 |
|
(29) |
Eriksson I, Sandback D, Ek B, Lindahl U, Kjellen L. cDNA cloning and sequencing of mouse mastocytoma glucosaminyl N-deacetylase/N-sulfotransferase, an enzyme involved in the biosynthesis of heparin. J. Biol. Chem. 269, 10438-10443, 1994 |
|
(30) |
Humphries DE, Lanciotti J, Karlinsky JB, cDNA cloning, genomic organization and chromosomal localization of human heparan glucosaminyl N-deacetylase/N-sulphotransferase-2. Biochem. J. 332, 303-307, 1998 |
|
(31) |
Aikawa J, Esko JD, Molecular cloning and expression of a third member of the heparan sulfate/heparin GlcNAc N-deacetylase/ N-sulfotransferase family. J. Biol. Chem. 274, 2690-2695, 1999 |
|
(32) |
Aikawa J, Grobe K, Tsujimoto M, Esko JD, Multiple isozymes of heparan sulfate/heparin GlcNAc N-deacetylase/GlcN N-sulfotransferase. Structure and activity of the fourth member, NDST4. J. Biol. Chem. 276, 5876-5882, 2001 |
|
(33) |
Kobayashi M, Habuchi H, Habuchi O, Saito M, Kimata K, Purification and characterization of heparan sulfate 2-sulfotransferase from cultured Chinese hamster ovary cells J. Biol. Chem. 271, 7645-7653, 1996 |
|
(34) |
Kobayashi M, Habuchi H, Yoneda M, Habuchi O, Kimata K, Molecular cloning and expression of Chinese hamster ovary cell heparan-sulfate 2-sulfotransferase. J. Biol. Chem. 272, 13980-13985, 1997 |
|
(35) |
Aquino RS, Landeira-Fernandez AM, Valente AP, Andrade LR, Mour o PA, Occurrence of sulfated galactans in marine angiosperms: evolutionary implications. Glycobiology. 15, 11-20, 2004 o PA, Occurrence of sulfated galactans in marine angiosperms: evolutionary implications. Glycobiology. 15, 11-20, 2004 |
|
(36) |
Kobayashi M, Sugumaran G, Liu J, Shworak NW, Silbert JE, Rosenberg RD, Molecular cloning and characterization of a human uronyl 2-sulfotransferase that sulfates iduronyl and glucuronyl residues in dermatan/chondroitin sulfate. J. Biol. Chem. 274, 10474-10480, 1999 |
|
(37) |
Ohtake S, Kimata K, Habuchi O, Recognition of sulfation pattern of chondroitin sulfate by uronosyl 2-O-sulfotransferase. J. Biol. Chem. 280, 39115-39123, 2005
|
|
(38) |
Habuchi H, Habuchi O, Kimata K, Purification and characterization of heparan sulfate 6-sulfotransferase from the culture medium of Chinese hamster ovary cells. J. Biol. Chem. 270, 4172-4179, 1995
|
|
(39) |
Habuchi H, Kobayashi M, Kimata K, Molecular characterization and expression of heparan-sulfate 6-sulfotransferase. Complete cDNA cloning in human and partial cloning in Chinese hamster ovary cells. J. Biol. Chem. 273, 9208-9213, 1998
|
|
(40) |
Habuchi H, Tanaka M, Habuchi O, Yoshida K, Suzuki H, Ban K, Kimata K, The occurrence of three isoforms of heparan sulfate 6-O-sulfotransferase having different specificities for hexuronic acid adjacent to the targeted N-sulfoglucosamine. J. Biol. Chem. 275, 2859-2868, 2000
|
|
(41) |
Habuchi H, Miyake G, Nogami K, Kuroiwa A, Matsuda Y, Kusche-Gullberg M, Habuchi O, Tanaka M, Kimata K, Biosynthesis of heparan sulphate with diverse structures and functions: two alternatively spliced forms of human heparan sulphate 6-O-sulphotransferase-2 having different expression patterns and properties. Biochem. J. 371,131-142, 2003
|
|
(42) |
Nagai N, Habuchi H., Esko JD, Kimata K, Stem domains of heparan sulfate 6-O-sulfotransferase are required for Golgi localization, oligomer formation and enzyme activity. J. Cell. Sci. 117, 3331-3341, 2004 |
|
(43) |
Liu J, Shworak NW, Fritze LM, Edelberg JM, Rosenberg RD, Purification of heparan sulfate D-glucosaminyl 3-O-sulfotransferase. J. Biol. Chem. 271, 27072-27082, 1996 |
|
(44) |
Shworak NW, Liu J, Fritze LM, Schwartz JJ, Zhang L, Logeart D, Rosenberg RD, Molecular cloning and expression of mouse and human cDNAs encoding heparan sulfate D-glucosaminyl 3-O-sulfotransferase. J. Biol. Chem. 272, 28008-28019, 1997
|
|
(45) |
Shworak NW, Liu J, Petros LM, Zhang L, Kobayashi M, Copeland NG, Jenkins NA, Rosenberg RD, Multiple isoforms of heparan sulfate D-glucosaminyl 3-O-sulfotransferase. Isolation, characterization, and expression of human cdnas and identification of distinct genomic loci. J. Biol. Chem. 274, 5170-5184, 1999
|
|
(46) |
Liu J, Shworak NW, Sinay P, Schwartz JJ, Zhang L, Fritze LM, Rosenberg RD, Expression of heparan sulfate D-glucosaminyl 3-O-sulfotransferase isoforms reveals novel substrate specificities. J. Biol. Chem. 274, 5185-5192, 1999
|
|
(47) |
Shukla D, Liu J, Blaiklock P, Shworak NW, Bai X, Esko JD, Cohen GH, Eisenberg RJ, Rosenberg RD, Spear PG, A novel role for 3-O-sulfated heparan sulfate in herpes simplex virus 1 entry. Cell. 99, 13-22, 1999
|
|
(48) |
Xia G, Chen J, Tiwari V, Ju W, Li JP, Malmstrom A, Shukla D, Liu J, Heparan sulfate 3-O-sulfotransferase isoform 5 generates both an antithrombin-binding site and an entry receptor for herpes simplex virus, type 1. Biol. Chem. 277, 37912-37919, 2002
|
|
(49) |
Xu D , Tiwari V, Xia G, Clement C, Shukla D, Liu J, Characterization of heparan sulphate 3-O-sulphotransferase isoform 6 and its role in assisting the entry of herpes simplex virus type 1. Biochem. J. 385, 451-459, 2005
|
|
(50) |
Uchimura K, Kadomatsu K, Nishimura H, Muramatsu H, Nakamura E, Kurosawa N, Habuchi O, El-Fasakhany M, Yoshikai Y, Muramatsu T, Functional analysis of the chondroitin 6-sulfotransferase gene in relation to lymphocyte subpopulations, brain development, and oversulfated chondroitin sulfates. J. Biol. Chem. 277, 1443-1450, 2002
|
|
(51) |
Kl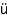 ppel M, Wight TN, Chan C, Hinek A, Wrana JL, Maintenance of chondroitin sulfation balance by chondroitin-4-sulfotransferase 1 is required for chondrocyte development and growth factor signaling during cartilage morphogenesis. Development, 132, 3989-4003, 2005 ppel M, Wight TN, Chan C, Hinek A, Wrana JL, Maintenance of chondroitin sulfation balance by chondroitin-4-sulfotransferase 1 is required for chondrocyte development and growth factor signaling during cartilage morphogenesis. Development, 132, 3989-4003, 2005
|
|
(52) |
Zhang H, Muramatsu T, Murase A, Yuasa S, Uchimura K, Kadomatsu K, N-Acetylglucosamine 6-O-sulfotransferase-1 is required for brain keratan sulfate biosynthesis and glial scar formation after brain injury. Glycobiology. 16, 702-710, 2006 |
|
(53) |
Ringvall M, Ledin J, Holmborn K, van Kuppevelt T, Ellin F, Eriksson I, Olofsson AM, Kjellen L, Forsberg E, Defective heparan sulfate biosynthesis and neonatal lethality in mice lacking N-deacetylase/N-sulfotransferase-1. J. Biol. Chem. 275, 25926-25930, 2000
|
|
(54) |
Fan G, Xiao L, Cheng L, Wang X, Sun B, Hu G, Targeted disruption of NDST-1 gene leads to pulmonary hypoplasia and neonatal respiratory distress in mice. FEBS Lett. 467, 7-11, 2000
|
|
(55) |
Grobe K, Inatani M, Pallerla SR, Castagnola J, Yamaguchi Y, Esko JD, Cerebral hypoplasia and craniofacial defects in mice lacking heparan sulfate Ndst1 gene function. Development. 132, 3777-3786, 2005
|
|
(56) |
Wang L, Fuster M, Sriramarao P, Esko JD, Endothelial heparan sulfate deficiency impairs L-selectin- and chemokine-mediated neutrophil trafficking during inflammatory responses. Nat. Immunol. 6, 902-910, 2005
|
|
(57) |
MacArthur JM, Bishop JR, Stanford KI, Wang L, Bensadoun A, Witztum JL, Esko JD, Liver heparan sulfate proteoglycans mediate clearance of triglyceride-rich lipoproteins independently of LDL receptor family members. J. Clin. Invest. 117, 153-64, 2007
|
|
(58) |
Humphries DE, Wong GW, Friend DS, Gurish MF, Qiu WT, Huang C, Sharpe AH, Stevens RL, Heparin is essential for the storage of specific granule proteases in mast cells. Nature. 400, 769-772, 1999
|
|
(59) |
Forsberg E, Pejler G, Ringvall M, Lunderius C, Tomasini-Johansson B, Kusche-Gullberg M, Eriksson I, Ledin J, Hellman L, Kjellen L, Abnormal mast cells in mice deficient in a heparin-synthesizing enzyme. Nature. 400, 773-776, 1999
|
|
(60) |
Bullock SL, Fletcher JM, Beddington RS, Wilson VA, Renal agenesis in mice homozygous for a gene trap mutation in the gene encoding heparan sulfate 2-sulfotransferase. Genes Dev. 12,1894-1906, 1998
|
|
(61) |
Habuchi H, Nagai N, Sugaya N, Atsumi F, Stevens RL, Kimata K, Mice deficient in heparan sulfate 6-O-sulfotransferase-1 exhibit defective heparan sulfate biosynthesis, abnormal placentation, and late embryonic lethality. J. Biol. Chem. 282, 15578-15588, 2007
|
|
(62) |
Pratt T, Conway CD, Tian NM, Price DJ, Mason JO, Heparan sulphation patterns generated by specific heparan sulfotransferase enzymes direct distinct aspects of retinal axon guidance at the optic chiasm. J. Neurosci. 26, 6911-6923, 2006
|
|
(63) |
HajMohammadi S, Enjyoji K, Princivalle M, Christi P, Lech M, Beeler D, Rayburn H, Schwartz JJ, Barzegar S, de Agostini AI, Post MJ, Rosenberg RD, Shworak NW, Normal levels of anticoagulant heparan sulfate are not essential for normal hemostasis. J. Clin. Invest. 111, 989?999, 2003
|
|
(64) |
Kobayashi T, Habuchi H, Tamura K, Ide H, Kimata K, Essential role of heparan sulfate 2-O-sulfotransferase in chick limb bud patterning and development. J. Biol. Chem. 282, 19589-19597, 2007
|
|
(65) |
Thiele H, Sakano M, Kitagawa H, Sugahara K, Rajab A, Hohne W, Ritter H, Leschik G, N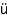 rnberg P, Mundlos S, Loss of chondroitin 6-O-sulfotransferase-1 function results in severe human chondrodysplasia with progressive spinal involvement. Proc. Natl. Acad. Sci. USA. 101, 10155-10160, 2004 rnberg P, Mundlos S, Loss of chondroitin 6-O-sulfotransferase-1 function results in severe human chondrodysplasia with progressive spinal involvement. Proc. Natl. Acad. Sci. USA. 101, 10155-10160, 2004
|
|
(66) |
Lin X, Perrimon N, Dally cooperates with Drosophila Frizzled 2 to transduce Wingless signalling. Nature. 400, 281-284, 1999
|
|
(67) |
Lin X, Buff EM, Perrimon N, Michelson AM, Heparan sulfate proteoglycans are essential for FGF receptor signaling during Drosophila embryonic development. Development. 126, 3715-3723, 1999
|
|
(68) |
Kamimura K, Fujise M, Villa F, Izumi S, Habuchi H, Kimata K, Nakato H, Drosophila heparan sulfate 6-O-sulfotransferase (dHS6ST) gene. Structure, expression, and function in the formation of the tracheal system. J. Biol. Chem. 276, 17014-17021, 2001
|
|
(69) |
Kamimura K, Koyama T, Habuchi H, Ueda R, Masu M, Kimata K, Nakato H, Specific and flexible roles of heparan sulfate modifications in Drosophila FGF signaling. J. Cell Biol. 174, 773-778, 2006
|
|
(70) |
Kamimura K, Rhodes JM, Ueda R, McNeely M, Shukla D, Kimata K, Spear PG, Shworak NW, Nakato H, Regulation of Notch signaling by Drosophila heparan sulfate 3-O sulfotransferase. J. Cell Biol. 166, 1069-1079, 2004
|
|
(71) |
Bulow HE, Hobert O, Differential sulfations and epimerization define heparan sulfate specificity in nervous system development. Neuron 41, 723-736, 2004
|
|
(72) |
Kinnunen T, Huang Z, Townsend J, Gatdula MM, Brown JR, Esko JD, Turnbull JE, Heparan 2-O-sulfotransferase, hst-2, is essential for normal cell migration in Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA. 102, 1507-1512, 2005
|
|
(73) |
Bink RJ, Habuchi H, Lele Z, Dolk E, Joore J, Rauch GJ, Geisler R, Wilson SW, den Hertog J, Kimata K, Zivkovic D, Heparan sulfate 6-o-sulfotransferase is essential for muscle development in zebrafish. J. Biol. Chem. 278, 31118-31127, 2003
|
|
(74) |
Chen E, Stringer SE, Rusch MA, Selleck SB, Ekker SC, A unique role for 6-O sulfation modification in zebrafish vascular development. Dev. Biol. 284, 364-376, 2005
|
|
(75) |
Merry CL, Bullock SL, Swan DC, Backen AC, Lyon M, Beddington RS, Wilson VA, Gallagher JT, The molecular phenotype of heparan sulfate in the Hs2st-/- mutant mouse. J. Biol. Chem. 276, 35429-35434, 2001
|
|
(76) |
Yusa A, Kitajima K, Habuchi O, N-linked oligosaccharides are required to produce and stabilize the active form of chondroitin 4-sulphotransferase-1. Biochem. J. 388, 115-121, 2005
|
|
(77) |
Yusa A, Kitajima K, Habuchi O, N-linked oligosaccharides on chondroitin 6-sulfotransferase-1 are required for production of the active enzyme, Golgi localization, and sulfotransferase activity toward keratan sulfate. J. Biol. Chem. 281, 20393-20403, 2006
|
|
(78) |
Uyama T, Ishida M, Izumikawa T, Trybala E, Tufaro F, Bergstrom T, Sugahara K, Kitagawa H, Chondroitin 4-O-sulfotransferase-1 regulates E disaccharide expression of chondroitin sulfate required for herpes simplex virus infectivity. J. Biol. Chem. 281, 38668-38674, 2006 |
|
(79) |
Habuchi O. Diversity and functions of glycosaminoglycan sulfotransferases. Biochim. Biophys. Acta, 1474, 115-127, 2000
|
|
(80) |
Kimata K, Habuchi O, Habuchi H, Watanabe H, Kockout Mice and Proteoglycans, Complehensive Glycoscience Vol. 4, Capter 3.11, Elsevier, 2007
|
| | |
| |